diff --git a/_toc.yml b/_toc.yml
index 121f6ba..99d5015 100644
--- a/_toc.yml
+++ b/_toc.yml
@@ -1,5 +1,4 @@
# Table of contents
-# Learn more at https://jupyterbook.org/customize/toc.html
format: jb-book
root: welcome
@@ -7,7 +6,8 @@ parts:
- caption: Introduction
chapters:
- file: introduction
-- caption: Getting Started
+ - file: prerequisites
+- caption: Quick Start
numbered: true
chapters:
- file: whole-game
@@ -23,12 +23,13 @@ parts:
- caption: Visualise
numbered: true
chapters:
+ - file: visualise
+ - file: vis-layers
- file: exploratory-data-analysis
- file: communicate-plots
- caption: Transform
numbered: true
chapters:
- - file: joins
- file: boolean-data
- file: numbers
- file: strings
@@ -36,13 +37,14 @@ parts:
- file: categorical-data
- file: dates-and-times
- file: missing-values
+ - file: joins
- caption: Import
numbered: true
chapters:
- file: spreadsheets
- - file: webscraping-and-apis
- - file: rectangling
- file: databases
+ - file: rectangling
+ - file: webscraping-and-apis
- caption: Programme
numbered: true
chapters:
diff --git a/categorical-data.ipynb b/categorical-data.ipynb
index e74ea07..40e9265 100644
--- a/categorical-data.ipynb
+++ b/categorical-data.ipynb
@@ -223,18 +223,17 @@
"source": [
"### Renaming Categories\n",
"\n",
- "Renaming categories is done by assigning new values to the `.cat.categories` property or by using the `rename_categories()` method (which works with a list or a dictionary)."
+ "Renaming categories is done via the `rename_categories()` method (which works with a list or a dictionary)."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "aedcf10f",
+ "id": "097171b8",
"metadata": {},
"outputs": [],
"source": [
- "df[\"cat_type\"].cat.categories = [\"alpha\", \"beta\", \"gamma\"]\n",
- "df"
+ "df[\"cat_type\"] = df[\"cat_type\"].cat.rename_categories([\"alpha\", \"beta\", \"gamma\"])"
]
},
{
@@ -380,7 +379,7 @@
"name": "python",
"nbconvert_exporter": "python",
"pygments_lexer": "ipython3",
- "version": "3.9.12"
+ "version": "3.10.12"
},
"toc-showtags": true
},
diff --git a/communicate-plots.ipynb b/communicate-plots.ipynb
index d351233..695ee37 100644
--- a/communicate-plots.ipynb
+++ b/communicate-plots.ipynb
@@ -12,26 +12,17 @@
"\n",
"In this chapter, you'll learn about using visualisation to communicate.\n",
"\n",
- "There are a plethora of options (and packages) for data visualisation using code. First, though a note about the different philosophies of data visualisation. There are broadly two categories of approach to using code to create data visualisations: imperative, where you build what you want, and declarative, where you say what you want. Choosing which to use involves a trade-off: imperative libraries offer you flexibility but at the cost of some verbosity; declarative libraries offer you a quick way to plot your data, but only if it’s in the right format to begin with, and customisation may be more difficult.\n",
- "\n",
- "There are also different purposes of data visualisation. It can be useful to bear in mind the three broad categories of visualisation that are out there:\n",
- "\n",
- "- exploratory\n",
- "\n",
- "- scientific\n",
- "\n",
- "- narrative\n",
- "\n",
- "Python has packages that cover all three of these.\n",
- "\n",
"In {ref}`exploratory-data-analysis`, you learned how to use plots as tools for *exploration*.\n",
- "When you make exploratory plots, you know---even before looking---which variables the plot will display.\n",
- "You made each plot for a purpose, could quickly look at it, and then move on to the next plot.\n",
- "In the course of most analyses, you'll produce tens or hundreds of plots, most of which are immediately thrown away. Exploratory visualisation is usually quick and dirty, and flexible too.\n",
+ "When you make exploratory plots, you know—even before looking—which variables the plot will display.\n",
+ "You made each plot for a purpose, quickly looked at it, and then moved on to the next plot.\n",
+ "In the course of most analyses, you'll produce tens or hundreds of plots, most of which are immediately thrown away.\n",
"\n",
- "The second kind, *scientific visualisation*, is the prime cut of your exploratory visualisation. It’s the kind of plot you might include in a more technical paper, the picture that says a thousand words. The first image of a black hole {cite}`akiyama2019first` is a prime example of this. You can get away with having a high density of information in a scientific plot because it's designed for specialists. Ensuring that important values can be accurately read from the plot is especially important in these kinds of charts. But they can also be the kind of plot that presents the killer results in a study; they might not be exciting to people who don’t look at charts for a living, but they might be exciting and, just as importantly, understandable by your peers.\n",
+ "Now that you understand your data, you need to *communicate* your understanding to others.\n",
+ "Your audience will likely not share your background knowledge and will not be deeply invested in the data. To help others quickly build up a good mental model of the data, you will need to invest considerable effort in making your plots as self-explanatory as possible. In this chapter, you'll learn some of the tools that **lets-plot** provides to do make charts tell a story.\n",
"\n",
- "The third and final kind is *narrative visualisation*, and it is the focus of this chapter—though we'll only scratch the surface. This is the one that requires the most thought in the step where you go from the first view to the end product because your audience will likely not share your background knowledge and will not be deeply invested in the data. It’s a visualisation that doesn’t just show a picture, but gives an insight. These are the kind of visualisations that you might see in the Financial Times, The Economist, or on the BBC News website. They come with aids that help the viewer focus on the aspects that the creator wanted them to (you can think of these aids or focuses as doing for visualisation what bold font does for text). They’re well worth using in your work, especially if you’re trying to communicate a particular narrative, and especially if the people you’re communicating with don’t have deep knowledge of the topic. You might use them in a paper that you hope will have a wide readership, in a blog post summarising your work, or in a report intended for a policymaker."
+ "### Prerequisities\n",
+ "\n",
+ "As ever, there are a plethora of options (and packages) for data visualisation using code. We're focusing on the declarative, \"grammar of graphics\" approach using **lets-plot** here, but advanced users looking for more complex graphics might wish to use an imperative library such as the excellent **matplotlib**. You should have both **lets-plot** and **pandas** installed. Once you have them installed, import them like so:"
]
},
{
@@ -54,24 +45,6 @@
"matplotlib_inline.backend_inline.set_matplotlib_formats(\"svg\")"
]
},
- {
- "cell_type": "markdown",
- "id": "17575f3a",
- "metadata": {},
- "source": [
- "### Prerequisites\n",
- "\n",
- "As well as **pandas**, you will need to install the declarative visualisation package **seaborn** for this chapter. This chapter uses the next generation version of **seaborn**, which can be installed by running the following on the command line (aka in the terminal): \n",
- "\n",
- "```bash\n",
- "pip install --pre seaborn\n",
- "```\n",
- "\n",
- "Although it will get installed when you install **seaborn**, we'll also be using the powerful imperative visualisation library that **seaborn** builds on, **matplotlib**.\n",
- "\n",
- "You'll need to import the **seaborn** and **pandas** libraries into your session using"
- ]
- },
{
"cell_type": "code",
"execution_count": null,
@@ -79,8 +52,12 @@
"metadata": {},
"outputs": [],
"source": [
- "import seaborn.objects as so\n",
- "import pandas as pd"
+ "from lets_plot import *\n",
+ "from lets_plot.mapping import as_discrete\n",
+ "import pandas as pd\n",
+ "import numpy as np\n",
+ "\n",
+ "LetsPlot.setup_html()"
]
},
{
@@ -88,9 +65,9 @@
"id": "a0dc9c10",
"metadata": {},
"source": [
- "## Labels and Titles\n",
+ "## Labels, titles, and other contextual information\n",
"\n",
- "The easiest place to start when turning an exploratory graphic into an expository graphic is with good labels. This example plot axis labels:"
+ "The easiest place to start when turning an exploratory graphic into an expository graphic is with good labels. Let's look at an example using the MPG (miles per gallon) data, which covers the fuel economy for 38 popular models of cars from 1999 to 2008."
]
},
{
@@ -111,21 +88,17 @@
"id": "1813ab08",
"metadata": {},
"source": [
- "Now let's do the plot with a title by passing the `title=` keyword argument into the `label` property."
+ "We want to show fuel efficiency on the highway changes with engine displacement, in litres. The most basic chart we can do with these variables is:"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "40b9bfcb",
+ "id": "c7574bc6",
"metadata": {},
"outputs": [],
"source": [
- "(\n",
- " so.Plot(mpg, x=\"displ\", y=\"hwy\")\n",
- " .add(so.Dot())\n",
- " .label(title=\"Fuel efficiency generally decreases with engine size\")\n",
- ")"
+ "(ggplot(mpg, aes(x=\"displ\", y=\"hwy\")) + geom_point())"
]
},
{
@@ -133,53 +106,114 @@
"id": "ff5ed0d4",
"metadata": {},
"source": [
- "The purpose of a plot title is to summarise the main finding. Avoid titles that just describe what the plot is, e.g. \"A scatterplot of engine displacement vs. fuel economy\".\n",
+ "Now we're going to add lots of extra useful information that will make the chart better. The purpose of a plot title is to summarize the main finding.\n",
+ "Avoid titles that just describe what the plot is, e.g., \"A scatterplot of engine displacement vs. fuel economy\".\n",
"\n",
- "If you need to add more text, there are two other useful labels that you can use:\n",
+ "We're going to:\n",
"\n",
- "- `subtitle` adds additional detail in a smaller font beneath the title.\n",
+ "- add a title that summarises the main finding you'd like the viewer to take away (as opposed to one just describing the obvious!)\n",
+ "- add a subtitle that provides more info on the y-axis, and make the x-label more understandable\n",
+ "- remove the y-axis label that is at an awkward viewing angle\n",
+ "- add a caption with the source of the data\n",
"\n",
- "- `caption` adds text at the bottom right of the plot, often used to describe the source of the data.\n"
+ "Putting this all in, we get:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "24b3513e",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\"))\n",
+ " + geom_point(aes(colour=\"class\"))\n",
+ " + geom_smooth(se=False, method=\"loess\", size=1)\n",
+ " + labs(\n",
+ " title=\"Fuel efficiency generally decreases with engine size\",\n",
+ " subtitle=\"Highway fuel efficiency (miles per gallon)\",\n",
+ " caption=\"Source: fueleconomy.gov\",\n",
+ " y=\"\",\n",
+ " x=\"Engine displacement (litres)\",\n",
+ " )\n",
+ ")"
]
},
{
"cell_type": "markdown",
- "id": "061f528b",
+ "id": "2e28877a",
"metadata": {},
"source": [
- "You can use `.label` to replace the axis and legend titles. It's usually a good idea to replace short variable names with more detailed descriptions, and to include the units."
+ "This is much clearer. It's easier to read, we know where the data come from, and we can see *why* we're being shown it too.\n",
+ "\n",
+ "But maybe we want a different message? You can flex depending on your needs, and some people prefer to have a rotated y-axis so that the subtitle can provide even more context:"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "b748ba52",
+ "id": "6489a6bf",
"metadata": {},
"outputs": [],
"source": [
"(\n",
- " so.Plot(mpg, x=\"displ\", y=\"hwy\")\n",
- " .add(so.Dot())\n",
- " .label(x=\"Engine displacement (L)\", y=\"Highway fuel economy (mpg)\")\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\"))\n",
+ " + geom_point(aes(colour=\"class\"))\n",
+ " + geom_smooth(se=False, method=\"loess\", size=1)\n",
+ " + labs(\n",
+ " x=\"Engine displacement (L)\",\n",
+ " y=\"Highway fuel economy (mpg)\",\n",
+ " colour=\"Car type\",\n",
+ " title=\"Fuel efficiency generally decreases with engine size\",\n",
+ " subtitle=\"Two seaters (sports cars) are an exception because of their light weight\",\n",
+ " caption=\"Source: fueleconomy.gov\",\n",
+ " )\n",
")"
]
},
{
"cell_type": "markdown",
- "id": "2e28877a",
+ "id": "9d88f188",
"metadata": {},
"source": [
- "It's possible to use mathematical equations and functions instead of text strings:"
+ "### Exercises\n",
+ "\n",
+ "1. Create one plot on the fuel economy data with customized `title`, `subtitle`, `caption`, `x`, `y`, and `color` labels.\n",
+ "\n",
+ "2. Recreate the following plot using the fuel economy data.\n",
+ " Note that both the colours and shapes of points vary by type of drive train."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "ffc13eab",
- "metadata": {},
+ "id": "683d547c",
+ "metadata": {
+ "tags": [
+ "remove-cell"
+ ]
+ },
"outputs": [],
"source": [
- "(so.Plot(mpg, x=\"displ\", y=\"hwy\").add(so.Dot()).label(y=str.capitalize, x=r\"$x^{y-z}$\"))"
+ "(\n",
+ " ggplot(mpg, aes(x=\"cty\", y=\"hwy\", color=\"drv\", shape=\"drv\"))\n",
+ " + geom_point()\n",
+ " + labs(\n",
+ " x=\"City MPG\",\n",
+ " y=\"Highway MPG\",\n",
+ " shape=\"Type of\\ndrive train\",\n",
+ " color=\"Type of\\ndrive train\",\n",
+ " )\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "e10cdbd9",
+ "metadata": {},
+ "source": [
+ "3. Take an exploratory graphic that you've created in the last month, and add informative titles to make it easier for others to understand."
]
},
{
@@ -189,65 +223,242 @@
"source": [
"## Annotations\n",
"\n",
- "[TODO]"
+ "In addition to labelling major components of your plot, it's often useful to label individual observations or groups of observations.\n",
+ "The first tool you have at your disposal is `geom_text()`.\n",
+ "`geom_text()` is similar to `geom_point()`, but it has an additional aesthetic: `label`.\n",
+ "This makes it possible to add textual labels to your plots.\n",
+ "\n",
+ "There are two possible sources of labels: ones that are part of the data, which we'll add with `geom_text`; and ones that we add directly and manually as annotations using `geom_label`.\n",
+ "\n",
+ "In the first case, you might have a dataframe that contains labels.\n",
+ "In the following plot we pull out the cars with the highest engine size in each drive type and save their information as a new data frame called `label_info`. In creating it, we pick out the mean values of \"hwy\" by \"drv\" as the points to label—but we could do any aggregation we feel would work well on the chart."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "60826a32",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "mapping = {\n",
+ " \"4\": \"4-wheel drive\",\n",
+ " \"f\": \"front-wheel drive\",\n",
+ " \"r\": \"rear-wheel drive\",\n",
+ "}\n",
+ "label_info = (\n",
+ " mpg.groupby(\"drv\")\n",
+ " .agg({\"hwy\": \"mean\", \"displ\": \"mean\"})\n",
+ " .reset_index()\n",
+ " .assign(drive_type=lambda x: x[\"drv\"].map(mapping))\n",
+ " .round(2)\n",
+ ")\n",
+ "label_info"
]
},
{
"cell_type": "markdown",
- "id": "1e245691",
+ "id": "93a136fe",
"metadata": {},
"source": [
- "## Scales\n",
- "\n",
- "The third way you can make your plot better for communication is to adjust the scales.\n",
- "Scales control the mapping from data values to things that you can perceive.\n",
- "Normally, **seaborn** automatically adds scales for you.\n",
- "For example, when you type:"
+ "Then, we use this new data frame to directly label the three groups to replace the legend with labels placed directly on the plot. Using the fontface and size arguments we can customize the look of the text labels. They’re larger than the rest of the text on the plot and bolded. (`theme(legend.position = \"none\")` turns all the legends off — we’ll talk about it more shortly.)"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "012d8c4d",
+ "id": "6f90c2aa",
"metadata": {},
"outputs": [],
"source": [
- "(so.Plot(mpg, x=\"displ\", y=\"hwy\", color=\"class\").add(so.Dot()))"
+ "(\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\", color=\"drv\"))\n",
+ " + geom_point(alpha=0.5)\n",
+ " + geom_smooth(se=False, method=\"loess\")\n",
+ " + geom_text(\n",
+ " aes(x=\"displ\", y=\"hwy\", label=\"drive_type\"),\n",
+ " data=label_info,\n",
+ " fontface=\"bold\",\n",
+ " size=8,\n",
+ " hjust=\"left\",\n",
+ " vjust=\"bottom\",\n",
+ " )\n",
+ " + theme(legend_position=\"none\")\n",
+ ")"
]
},
{
"cell_type": "markdown",
- "id": "fae05382",
+ "id": "98c17829",
"metadata": {},
"source": [
- "**seaborn** automatically adds default scales behind the scenes:"
+ "Note the use of `hjust` (horizontal justification) and `vjust` (vertical justification) to control the alignment of the label.\n",
+ "\n",
+ "\n",
+ "The second of the two methods we're looking at is `geom_label`. This has two modes: in the first, it works like `geom_text` but with a box around the text, like so:"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "f1b10368",
+ "id": "bdcd79bb",
"metadata": {},
"outputs": [],
"source": [
+ "potential_outliers = mpg.query(\"hwy > 40 | (hwy > 20 & displ > 5)\")\n",
"(\n",
- " so.Plot(mpg, x=\"displ\", y=\"hwy\", color=\"class\")\n",
- " .add(so.Dot())\n",
- " .scale(\n",
- " x=so.Continuous(),\n",
- " y=so.Continuous(),\n",
- " color=so.Nominal(),\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\"))\n",
+ " + geom_point(color=\"black\")\n",
+ " + geom_smooth(se=False, method=\"loess\", color=\"black\")\n",
+ " + geom_point(\n",
+ " data=potential_outliers,\n",
+ " color=\"red\",\n",
" )\n",
+ " + geom_label(\n",
+ " aes(label=\"model\"),\n",
+ " data=potential_outliers,\n",
+ " color=\"red\",\n",
+ " position=position_jitter(),\n",
+ " fontface=\"bold\",\n",
+ " size=5,\n",
+ " hjust=\"left\",\n",
+ " vjust=\"bottom\",\n",
+ " )\n",
+ " + theme(legend_position=\"none\")\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "38b69dcf",
+ "metadata": {},
+ "source": [
+ "The second method is generally useful for adding either a single or several annotations to a plot, like so:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "d1e2cc3a",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import textwrap\n",
+ "\n",
+ "# wrap the text so it is over multiple lines:\n",
+ "trend_text = textwrap.fill(\"Larger engine sizes tend to have lower fuel economy.\", 30)\n",
+ "trend_text"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "e8c09f57",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\"))\n",
+ " + geom_point()\n",
+ " + geom_label(x=3.5, y=38, label=trend_text, hjust=\"left\", color=\"red\")\n",
+ " + geom_segment(x=2, y=40, xend=5, yend=25, arrow=arrow(type=\"closed\"), color=\"red\")\n",
")"
]
},
{
"cell_type": "markdown",
- "id": "03b4d70c",
+ "id": "0720e7eb",
+ "metadata": {},
+ "source": [
+ "Annotation is a powerful tool for communicating main takeaways and interesting features of your visualisations. The only limit is your imagination (and your patience with positioning annotations to be aesthetically pleasing)!"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "9c00a0fd",
+ "metadata": {},
+ "source": [
+ "Remember, in addition to `geom_text()` and `geom_label()`, you have many other geoms in **lets-plot** available to help annotate your plot.\n",
+ "A couple ideas:\n",
+ "\n",
+ "- Use `geom_hline()` and `geom_vline()` to add reference lines.\n",
+ " We often make them thick (`size = 2`) and grey (`color = gray`), and draw them underneath the primary data layer.\n",
+ " That makes them easy to see, without drawing attention away from the data.\n",
+ "\n",
+ "- Use `geom_rect()` to draw a rectangle around points of interest.\n",
+ " The boundaries of the rectangle are defined by aesthetics `xmin`, `xmax`, `ymin`, `ymax`.\n",
+ "\n",
+ "- You already saw the use of `geom_segment()` with the `arrow` argument to draw attention to a point with an arrow.\n",
+ " Use aesthetics `x` and `y` to define the starting location, and `xend` and `yend` to define the end location.\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "730162e6",
+ "metadata": {},
+ "source": [
+ "### Exercises\n",
+ "\n",
+ "1. Use `geom_text()` with infinite positions to place text at the four corners of the plot.\n",
+ "\n",
+ "2. Use `geom_label()` to add a point geom in the middle of your last plot without having to create a dataframe\n",
+ " Customise the shape, size, or colour of the point.\n",
+ "\n",
+ "3. How do labels with `geom_text()` interact with faceting?\n",
+ " How can you add a label to a single facet?\n",
+ " How can you put a different label in each facet?\n",
+ " (Hint: Think about the dataset that is being passed to `geom_text()`.)\n",
+ "\n",
+ "4. What arguments to `geom_label()` control the appearance of the background box?\n",
+ "\n",
+ "5. What are the four arguments to `arrow()`?\n",
+ " How do they work?\n",
+ " Create a series of plots that demonstrate the most important options.\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "2f665492",
+ "metadata": {},
+ "source": [
+ "## Scales\n",
+ "\n",
+ "Another you can make your plot better for communication is to adjust the scales.\n",
+ "Scales control how the aesthetic mappings manifest visually.\n",
+ "\n",
+ "### Default scales\n",
+ "\n",
+ "Normally, **lets-plot** automatically adds scales for you and you don't need to worry about them. For example, when you type:\n",
+ "\n",
+ "```python\n",
+ "(\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\")) +\n",
+ " geom_point(aes(color=\"class\"))\n",
+ ")\n",
+ "```\n",
+ "\n",
+ "**lets-plot** is automatically doing this behind the scenes:\n",
+ "\n",
+ "```python\n",
+ "(\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\")) +\n",
+ " geom_point(aes(color=\"class\")) +\n",
+ " scale_x_continous() +\n",
+ " scale_y_continuous() +\n",
+ " scale_color_discrete()\n",
+ ")\n",
+ "```"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "39332b3b",
"metadata": {},
"source": [
- "Note the naming scheme for scales: `.scale` followed by the name of the dimension, then `=so.`, then the name of the scale.\n",
- "The default scales are named according to the type of variable they align with: continuous, nominal, and so on.\n",
+ "Note the naming scheme for scales: `scale_` followed by the name of the aesthetic, then `_`, then the name of the scale.\n",
+ "The default scales are named according to the type of variable they align with: continuous, discrete, datetime, or date.\n",
+ "`scale_x_continuous()` puts the numeric values from `displ` on a continuous number line on the x-axis, `scale_color_discrete()` chooses colours for each of the `class` of car, etc.\n",
+ "There are lots of non-default scales which you'll learn about below.\n",
"\n",
"The default scales have been carefully chosen to do a good job for a wide range of inputs.\n",
"Nevertheless, you might want to override the defaults for two reasons:\n",
@@ -256,195 +467,843 @@
" This allows you to do things like change the breaks on the axes, or the key labels on the legend.\n",
"\n",
"- You might want to replace the scale altogether, and use a completely different algorithm.\n",
- " Often you can do better than the default because you know more about the data.\n",
+ " Often you can do better than the default because you know more about the data.\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "c645247f",
+ "metadata": {},
+ "source": [
+ "### Axis ticks and legend keys\n",
"\n",
- "```{admonition} Exercise\n",
- "Try a plot with a scale setting of `x=\"log\"`.\n",
- "```"
+ "Collectively axes and legends get the somewhat confusing name **guides** in **lets-plot**. Axes are used for x and y aesthetics; legends are used for everything else.\n",
+ "\n",
+ "There are two primary arguments that affect the appearance of the ticks on the axes and the keys on the legend: `breaks` and `labels`.\n",
+ "Breaks controls the position of the ticks, or the values associated with the keys. If you like, the breaks *are* the ticks.\n",
+ "Labels controls the text label associated with each tick/key. We might more accurately call these *tick labels*.\n",
+ "The most common use of `breaks` is to override the default choice:\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "a95604d8",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\", color=\"drv\"))\n",
+ " + geom_point()\n",
+ " + scale_y_continuous(breaks=np.arange(15, 40, step=5))\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "bd1113b7",
+ "metadata": {},
+ "source": [
+ "You can use `labels` in the same way (ie pass in an array or list of strings the same length as `breaks`). To remove them altogether, you would have to use a theme, though, a topic we'll return to later.\n",
+ "You can also use `breaks` and `labels` to control the appearance of legends.\n",
+ "For discrete scales for categorical variables, `labels` can be a named list of the existing levels names and the desired labels for them.\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "1a852304",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\", color=\"drv\"))\n",
+ " + geom_point()\n",
+ " + scale_color_discrete(labels=[\"4-wheel\", \"front\", \"rear\"])\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "184dfb30",
+ "metadata": {},
+ "source": [
+ "To change the formatting of the tick labels, use the `format=` keyword argument. This is useful to render currencies, percentages, and so on—though it's often easier for the reader to just see this symbol once in the axis label.\n",
+ "\n",
+ "In the example below, we read in the `diamonds` dataset and then format it with a command `format=\"$.2s\"`; let's break this down:\n",
+ "\n",
+ "- the dollar sign says put a dollar sign in front of every number\n",
+ "- the .2 says use two significant digits\n",
+ "- the s says, use the Système International (SI)\n",
+ "\n",
+ "There are a wealth of alternative options for formatting—it's best to use the [helpful page on formatting](https://lets-plot.org/pages/formats.html) in the documentation of **lets-plot** to find out more."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "40ac230e",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "diamonds = pd.read_csv(\n",
+ " \"https://vincentarelbundock.github.io/Rdatasets/csv/ggplot2/diamonds.csv\",\n",
+ " index_col=0,\n",
+ ")\n",
+ "diamonds[\"cut\"] = diamonds[\"cut\"].astype(\n",
+ " pd.CategoricalDtype(\n",
+ " categories=[\"Fair\", \"Good\", \"Very Good\", \"Premium\", \"Ideal\"], ordered=True\n",
+ " )\n",
+ ")\n",
+ "diamonds[\"color\"] = diamonds[\"color\"].astype(\n",
+ " pd.CategoricalDtype(categories=[\"D\", \"E\", \"F\", \"G\", \"H\", \"I\", \"J\"], ordered=True)\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "1520bb3c",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(\n",
+ " ggplot(diamonds, aes(x=\"cut\", y=\"price\"))\n",
+ " + geom_boxplot()\n",
+ " + coord_flip()\n",
+ " + scale_y_continuous(format=\"$.2s\", breaks=np.arange(0, 19000, step=6000))\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "6f2edc1b",
+ "metadata": {},
+ "source": [
+ "Another use of breaks is when you have relatively few data points and want to highlight exactly where the observations occur. For example, take this plot that shows when each US president started and ended their term."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "9d1f993a",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "presidential = pd.read_csv(\n",
+ " \"https://vincentarelbundock.github.io/Rdatasets/csv/ggplot2/presidential.csv\",\n",
+ " index_col=0,\n",
+ ")\n",
+ "presidential = presidential.astype({\"start\": \"datetime64[ns]\", \"end\": \"datetime64[ns]\"})\n",
+ "presidential[\"id\"] = 33 + presidential.index\n",
+ "presidential.head()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "cd2cc430",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# (\n",
+ "# ggplot(presidential, aes(x=\"start\", y=\"id\")) +\n",
+ "# geom_point() +\n",
+ "# geom_segment(aes(xend=\"end\", yend=\"id\")) +\n",
+ "# scale_x_datetime(breaks=presidential[\"start\"])\n",
+ "# )"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "a2d0b3f1",
+ "metadata": {},
+ "source": [
+ "Note that this example isn't currently working; we have [raised an issue on the **letsplot** Github page](https://github.com/JetBrains/lets-plot/issues/346) to get it fixed.\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "8b451c76",
+ "metadata": {},
+ "source": [
+ "### Legend layout\n",
+ "\n",
+ "You will most often use `breaks` and `labels` to tweak the axes.\n",
+ "While they both also work for legends, there are a few other techniques you are more likely to use.\n",
+ "\n",
+ "To control the overall position of the legend, you need to use a `theme()` setting.\n",
+ "We'll come back to themes at the end of the chapter, but in brief, they control the non-data parts of the plot.\n",
+ "The theme setting `legend.position` controls where the legend is drawn, and to demonstrate this we'll use `gggrid` to arrange all of the plots."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "52d6e86a",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "base = ggplot(mpg, aes(x=\"displ\", y=\"hwy\")) + geom_point(aes(color=\"class\"))\n",
+ "\n",
+ "p1 = base + theme(legend_position=\"right\") # the default\n",
+ "p2 = base + theme(legend_position=\"left\")\n",
+ "p3 = base + theme(legend_position=\"top\") + guides(color=guide_legend(nrow=3))\n",
+ "p4 = base + theme(legend_position=\"bottom\") + guides(color=guide_legend(nrow=3))\n",
+ "\n",
+ "gggrid([p1, p2, p3, p4], ncol=2)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "7ce2507b",
+ "metadata": {},
+ "source": [
+ "If your plot is short and wide, place the legend at the top or bottom, and if it's tall and narrow, place the legend at the left or right. You can also use `legend_position = \"none\"` to suppress the display of the legend altogether.\n",
+ "\n",
+ "To control the display of individual legends, use `guides()` along with `guide_legend()` or `guide_colorbar()`."
]
},
{
"cell_type": "markdown",
- "id": "6845649b",
+ "id": "f27913c7",
"metadata": {},
"source": [
- "### Axis Ticks\n",
"\n",
- "You can specify axis ticks directly using the `tick` property on the `Scale` parameter:"
+ "### Replacing a scale\n",
+ "\n",
+ "Instead of just tweaking the details a little, you can instead replace the scale altogether.\n",
+ "There are two types of scales you're mostly likely to want to switch out: continuous position scales and colour scales.\n",
+ "Fortunately, the same principles apply to all the other aesthetics, so once you've mastered position and colour, you'll be able to quickly pick up other scale replacements.\n",
+ "\n",
+ "It's very useful to plot transformations of your variable.\n",
+ "For example, it's easier to see the precise relationship between `carat` and `price` if we log transform them. The way to do this is by using an `apply` function on the data that gets sent to `ggplot`:"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "dcd15c76",
+ "id": "2c1d3f8d",
"metadata": {},
"outputs": [],
"source": [
"(\n",
- " so.Plot(mpg, x=\"displ\", y=\"hwy\", color=\"class\")\n",
- " .add(so.Dot())\n",
- " .scale(\n",
- " x=so.Continuous(),\n",
- " y=so.Continuous().tick(at=[0, 10, 20, 30, 40]),\n",
- " color=so.Nominal(),\n",
+ " ggplot(\n",
+ " diamonds.apply({\"carat\": np.log10, \"price\": np.log10}),\n",
+ " aes(x=\"carat\", y=\"price\"),\n",
" )\n",
+ " + geom_bin2d()\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "f19dbbec",
+ "metadata": {},
+ "source": [
+ "However, the disadvantage of this transformation is that the axes are now mislabelled with the original values, making it hard to interpret the plot. Instead of doing the transformation in the aesthetic mapping, we can instead do it with the scale. This is visually identical, except the axes are labelled on the original data scale."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "39b4ef8d",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(\n",
+ " ggplot(diamonds, aes(x=\"carat\", y=\"price\"))\n",
+ " + geom_bin2d()\n",
+ " + scale_x_log10()\n",
+ " + scale_y_log10()\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "4402c4de",
+ "metadata": {},
+ "source": [
+ "Another scale that is frequently customised is colour. The default categorical scale picks colors that are evenly spaced around the color wheel. Useful alternatives are the ColorBrewer scales which have been hand tuned to work better for people with common types of colour blindness. The two plots below look similar, but there is enough difference in the shades of red and green that the dots on the right can be distinguished even by people with red-green colour blindness."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "f06d7e40",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"displ\", y=\"hwy\")) + geom_point(aes(color=\"drv\")))"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "6186b520",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\"))\n",
+ " + geom_point(aes(color=\"drv\"))\n",
+ " + scale_color_brewer(palette=\"Set1\")\n",
")"
]
},
{
"cell_type": "markdown",
- "id": "f56d0216",
+ "id": "f19af6ad",
"metadata": {},
"source": [
- "### Legend Keys"
+ "Don't forget simpler techniques for improving accessibility.\n",
+ "If there are just a few colors, you can add a redundant shape mapping.\n",
+ "This will also help ensure your plot is interpretable in black and white."
]
},
{
"cell_type": "markdown",
- "id": "d3631634",
+ "id": "253af5a4",
"metadata": {},
"source": [
- "### Legend Layout\n",
+ "The ColorBrewer scales are documented online at . The sequential (top) and diverging (bottom) palettes are particularly useful if your categorical values are ordered, or have a \"middle\". This often arises if you've used `pd.cut()` to make a continuous variable into a categorical variable."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "bd347524",
+ "metadata": {
+ "tags": [
+ "remove-input"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "# remove-input\n",
+ "cmaps = [\n",
+ " (\n",
+ " \"Perceptually Uniform Sequential\",\n",
+ " [\"viridis\", \"plasma\", \"inferno\", \"magma\", \"cividis\"],\n",
+ " ),\n",
+ " (\n",
+ " \"Sequential\",\n",
+ " [\n",
+ " \"Blues\",\n",
+ " \"BuGn\",\n",
+ " \"BuPu\",\n",
+ " \"GnBu\",\n",
+ " \"Greens\",\n",
+ " \"Greys\",\n",
+ " \"Oranges\",\n",
+ " \"OrRd\",\n",
+ " \"PuBu\",\n",
+ " \"PuBuGn\",\n",
+ " \"PuRd\",\n",
+ " \"Purples\",\n",
+ " \"RdPu\",\n",
+ " \"Reds\",\n",
+ " \"YlGn\",\n",
+ " \"YlGnBu\",\n",
+ " \"YlOrBr\",\n",
+ " \"YlOrRd\",\n",
+ " ],\n",
+ " ),\n",
+ " (\n",
+ " \"Diverging\",\n",
+ " [\n",
+ " \"BrBG\",\n",
+ " \"PiYG\",\n",
+ " \"PRGn\",\n",
+ " \"PuOr\",\n",
+ " \"RdBu\",\n",
+ " \"RdGy\",\n",
+ " \"RdYlBu\",\n",
+ " \"RdYlGn\",\n",
+ " ],\n",
+ " ),\n",
+ " (\n",
+ " \"Qualitative\",\n",
+ " [\n",
+ " \"Pastel1\",\n",
+ " \"Pastel2\",\n",
+ " \"Paired\",\n",
+ " \"Accent\",\n",
+ " \"Dark2\",\n",
+ " \"Set1\",\n",
+ " \"Set2\",\n",
+ " \"Set3\",\n",
+ " \"tab10\",\n",
+ " \"tab20\",\n",
+ " \"tab20b\",\n",
+ " \"tab20c\",\n",
+ " ],\n",
+ " ),\n",
+ "]\n",
+ "\n",
+ "\n",
+ "gradient = np.linspace(0, 1, 256)\n",
+ "gradient = np.vstack((gradient, gradient))\n",
"\n",
- "[TODO]"
+ "\n",
+ "def plot_color_gradients(cmap_category, cmap_list):\n",
+ " # Create figure and adjust figure height to number of colourmaps\n",
+ " nrows = len(cmap_list)\n",
+ " figh = 0.35 + 0.15 + (nrows + (nrows - 1) * 0.1) * 0.22\n",
+ " fig, axs = plt.subplots(nrows=nrows, figsize=(6.4, figh))\n",
+ " fig.subplots_adjust(top=1 - 0.35 / figh, bottom=0.15 / figh, left=0.2, right=0.99)\n",
+ "\n",
+ " axs[0].set_title(cmap_category + \" colormaps\", fontsize=14)\n",
+ "\n",
+ " for ax, name in zip(axs, cmap_list):\n",
+ " ax.imshow(gradient, aspect=\"auto\", cmap=plt.get_cmap(name))\n",
+ " ax.text(\n",
+ " -0.01,\n",
+ " 0.5,\n",
+ " name,\n",
+ " va=\"center\",\n",
+ " ha=\"right\",\n",
+ " fontsize=10,\n",
+ " transform=ax.transAxes,\n",
+ " )\n",
+ "\n",
+ " # Turn off *all* ticks & spines, not just the ones with colourmaps.\n",
+ " for ax in axs:\n",
+ " ax.set_axis_off()\n",
+ "\n",
+ "\n",
+ "for cmap_category, cmap_list in cmaps[1:2]:\n",
+ " plot_color_gradients(cmap_category, cmap_list)\n",
+ "\n",
+ "plt.show()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "d6350c71",
+ "metadata": {
+ "tags": [
+ "remove-input"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "# remove input\n",
+ "for cmap_category, cmap_list in cmaps[3:4]:\n",
+ " plot_color_gradients(cmap_category, cmap_list)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "0063a574",
+ "metadata": {
+ "tags": [
+ "remove-input"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "# remove input\n",
+ "for cmap_category, cmap_list in cmaps[2:3]:\n",
+ " plot_color_gradients(cmap_category, cmap_list)"
]
},
{
"cell_type": "markdown",
- "id": "57d59667",
+ "id": "c32c2237",
+ "metadata": {},
+ "source": [
+ "When you have a predefined mapping between values and colours, use `scale_color_manual()`. For example, if we map presidential party to colour, we want to use the standard mapping of red for Republicans and blue for Democrats. One approach for assigning these colors is using hex colour codes:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "9751058d",
"metadata": {},
+ "outputs": [],
"source": [
- "### Limits, aka 'zooming'\n",
+ "mini_presid = presidential.iloc[5:, :]\n",
"\n",
- "There are two ways to control the plot limits:\n",
+ "(\n",
+ " ggplot(mini_presid, aes(x=\"start\", y=\"id\", color=\"party\"))\n",
+ " + geom_point(size=3)\n",
+ " + geom_segment(aes(xend=\"end\", yend=\"id\"), size=1)\n",
+ " + scale_x_datetime(breaks=mini_presid[\"start\"], format=\"%Y\")\n",
+ " + scale_color_manual(values=[\"#00AEF3\", \"#E81B23\"], name=\"party\")\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "6be370b4",
+ "metadata": {},
+ "source": [
+ "You can also use typical colour names such as \"red\" and \"blue\".\n",
+ "\n",
+ "For continuous colour, you can use the built-in `scale_color_gradient()` or `scale_fill_gradient()`.\n",
+ "If you have a diverging scale, you can use `scale_color_gradient2()`. That allows you to give, for example, positive and negative values different colors. That's sometimes also useful if you want to distinguish points above or below the mean.\n",
"\n",
- "1. Adjusting what data are plotted\n",
- "2. Setting the limits in each scale\n",
+ "Another option is to use the viridis, magma, inferno, and plasma color scales developed for the extremely powerful imperative Python plotting package **[matplotlib](https://matplotlib.org/)**. The designers, Nathaniel Smith and Stéfan van der Walt, carefully tailored continuous color schemes that are perceptible to people with various forms of color blindness as well as perceptually uniform in both color and black and white. These scales are available as palettes in *lets-plot*. Here's an example using the continuous version of viridis (we'll generate some random data first):"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "644fd814",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "prng = np.random.default_rng(1837) # prng=probabilistic random number generator\n",
+ "df_rnd = pd.DataFrame(prng.standard_normal((1000, 2)), columns=[\"x\", \"y\"])\n",
+ "(\n",
+ " ggplot(df_rnd, aes(x=\"x\", y=\"y\"))\n",
+ " + geom_bin2d()\n",
+ " + coord_fixed()\n",
+ " + scale_fill_viridis(option=\"plasma\")\n",
+ " + labs(title=\"Plasma, continuous\")\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "e7cf0931",
+ "metadata": {},
+ "source": [
+ "### Zooming\n",
+ "\n",
+ "There are three ways to control the plot limits:\n",
+ "\n",
+ "1. Adjusting what data are plotted.\n",
+ "2. Setting the limits in each scale.\n",
+ "3. Setting `xlim` and `ylim` in `coord_cartesian()`.\n",
+ "\n",
+ "We'll demonstrate these options in a series of plots.\n",
+ "The first plot shows the relationship between engine size and fuel efficiency, coloured by type of drive train.\n",
+ "The second plot shows the same variables, but subsets the data that are plotted.\n",
+ "Subsetting the data has affected the x and y scales as well as the smooth curve.\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "25a29f38",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\"))\n",
+ " + geom_point(aes(color=\"drv\"))\n",
+ " + geom_smooth(method=\"loess\")\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "42318a59",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "mpg_condition = (\n",
+ " (mpg[\"displ\"] >= 5) & (mpg[\"displ\"] <= 6) & (mpg[\"hwy\"] >= 10) & (mpg[\"hwy\"] <= 25)\n",
+ ")\n",
"\n",
- "Here is the same plot done according to 1 and 2 respectively."
+ "(\n",
+ " ggplot(mpg.loc[mpg_condition], aes(x=\"displ\", y=\"hwy\"))\n",
+ " + geom_point(aes(color=\"drv\"))\n",
+ " + geom_smooth(method=\"loess\")\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "ec4c07d0",
+ "metadata": {},
+ "source": [
+ "Let's compare these to the two plots below where the first plot sets the `limits` on individual scales and the second plot sets them in `coord_cartesian()`.\n",
+ "We can see that reducing the limits is equivalent to subsetting the data.\n",
+ "Therefore, to zoom in on a region of the plot, it's generally best to use `coord_cartesian()`."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "013471ab",
+ "id": "03001d5e",
"metadata": {},
"outputs": [],
"source": [
"(\n",
- " so.Plot(mpg, x=\"displ\", y=\"hwy\", color=\"class\")\n",
- " .add(so.Dot())\n",
- " .limit(x=(5, 7), y=(10, 30))\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\"))\n",
+ " + geom_point(aes(color=\"drv\"))\n",
+ " + geom_smooth(method=\"loess\")\n",
+ " + scale_x_continuous(limits=(5, 6))\n",
+ " + scale_y_continuous(limits=(10, 25))\n",
")"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "1734b4f4",
+ "id": "dc3bb833",
"metadata": {},
"outputs": [],
"source": [
"(\n",
- " so.Plot(\n",
- " mpg.query(\"displ >= 5 & displ <= 7 & hwy >= 10 & hwy <= 30\"),\n",
- " x=\"displ\",\n",
- " y=\"hwy\",\n",
- " color=\"class\",\n",
- " ).add(so.Dot())\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\"))\n",
+ " + geom_point(aes(color=\"drv\"))\n",
+ " + geom_smooth(method=\"loess\")\n",
+ " + coord_cartesian(xlim=(5, 6), ylim=(10, 25))\n",
")"
]
},
{
"cell_type": "markdown",
- "id": "89ad530b",
+ "id": "5d1fc3ee",
+ "metadata": {},
+ "source": [
+ "On the other hand, setting the `limits` on individual scales is generally more useful if you want to *expand* the limits, e.g., to match scales across different plots.\n",
+ "For example, if we extract two classes of cars and plot them separately, it's difficult to compare the plots because all three scales (the x-axis, the y-axis, and the colour aesthetic) have different ranges."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "aee538a8",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "suv = mpg.loc[mpg[\"class\"] == \"suv\"]\n",
+ "compact = mpg.loc[mpg[\"class\"] == \"compact\"]\n",
+ "(ggplot(suv, aes(x=\"displ\", y=\"hwy\", color=\"drv\")) + geom_point())"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "a82c8c23",
"metadata": {},
+ "outputs": [],
"source": [
- "While they convey the same information, the former looks better."
+ "(ggplot(compact, aes(x=\"displ\", y=\"hwy\", color=\"drv\")) + geom_point())"
]
},
{
"cell_type": "markdown",
- "id": "eeaa7fde",
+ "id": "be777179",
+ "metadata": {},
+ "source": [
+ "One way to overcome this problem is to share scales across multiple plots, training the scales with the `limits` of the full data.\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "db6fce43",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "x_scale = scale_x_continuous(limits=mpg[\"displ\"].agg([\"max\", \"min\"]).tolist())\n",
+ "y_scale = scale_y_continuous(limits=mpg[\"hwy\"].agg([\"max\", \"min\"]).tolist())\n",
+ "col_scale = scale_color_discrete(limits=mpg[\"drv\"].unique())"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "dd9e6606",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(\n",
+ " ggplot(suv, aes(x=\"displ\", y=\"hwy\", color=\"drv\"))\n",
+ " + geom_point()\n",
+ " + x_scale\n",
+ " + y_scale\n",
+ " + col_scale\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "bdd8b2c5",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(\n",
+ " ggplot(compact, aes(x=\"displ\", y=\"hwy\", color=\"drv\"))\n",
+ " + geom_point()\n",
+ " + x_scale\n",
+ " + y_scale\n",
+ " + col_scale\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "577d8648",
+ "metadata": {},
+ "source": [
+ "In this particular case, you could have simply used faceting, but this technique is useful more generally, if for instance, you want to spread plots over multiple pages of a report.\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "4094830b",
+ "metadata": {},
+ "source": [
+ "### Exercises\n",
+ "\n",
+ "1. What is the first argument to every scale?\n",
+ " How does it compare to `labs()`?\n",
+ "\n",
+ "2. Change the display of the presidential terms by:\n",
+ "\n",
+ " a. Combining the two variants that customize colors and x axis breaks.\n",
+ " b. Improving the display of the y axis.\n",
+ " c. Labelling each term with the name of the president.\n",
+ " d. Adding informative plot labels.\n",
+ " e. Placing breaks every 4 years (this is trickier than it seems!).\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "8b574471",
"metadata": {},
"source": [
"## Themes\n",
"\n",
- "Seaborn comes with several built-in themes that you can switch between by using"
+ "Finally, you can customise the non-data elements of your plot with a theme:\n"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "8749bccd",
+ "id": "0b2364ca",
"metadata": {},
"outputs": [],
"source": [
- "import seaborn as sns\n",
- "\n",
- "sns.set_theme(style=\"darkgrid\", palette=\"dark\")\n",
+ "(\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\"))\n",
+ " + geom_point(aes(color=\"class\"))\n",
+ " + geom_smooth(se=False)\n",
+ " + theme_grey()\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "7814bb4d",
+ "metadata": {},
+ "source": [
+ "**lets-plot** includes several built-in themes that you can find [here](https://lets-plot.org/pages/api.html#predefined-themes). You can also create your own themes, if you are trying to match a particular corporate or journal style.\n",
"\n",
- "(so.Plot(mpg, x=\"displ\", y=\"hwy\", color=\"class\").add(so.Dot()))"
+ "Here's an example of changing multiple `theme()` settings:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "67bfa9c8",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(\n",
+ " ggplot(mpg, aes(x=\"displ\", color=\"drv\"))\n",
+ " + geom_density(size=2)\n",
+ " + ggtitle(\"Density of drives\")\n",
+ " + theme(\n",
+ " axis_line=element_line(size=4),\n",
+ " axis_ticks_length=10,\n",
+ " axis_title_y=\"blank\",\n",
+ " legend_position=[1, 1],\n",
+ " legend_justification=[1, 1],\n",
+ " panel_background=element_rect(color=\"black\", fill=\"#eeeeee\", size=2),\n",
+ " panel_grid=element_line(color=\"black\", size=1),\n",
+ " )\n",
+ ")"
]
},
{
"cell_type": "markdown",
- "id": "b09cf2b2",
+ "id": "5b05b5da",
"metadata": {},
"source": [
- "Note that you can also create your own themes using **matplotlib**, the library that sits under **seaborn** (this book uses a custom theme).\n"
+ "### Exercises\n",
+ "\n",
+ "1. Make the axis labels of your plot blue and bolded.\n"
]
},
{
"cell_type": "markdown",
- "id": "310e4b73",
+ "id": "a56216db",
"metadata": {},
"source": [
- "## Saving Plots\n",
+ "## Layout\n",
"\n",
- "There are lots of output options to choose from to save your file to. Remember that, for graphics, *vector formats* are generally better than *raster formats*. In practice, this means saving plots in svg or pdf formats over jpg or png file formats. The svg format works in a lot of contexts (including Microsoft Word) and is a good default. To choose between formats, just supply the file extension and the file type will change automatically, eg \"chart.svg\" for svg or \"chart.png\" for png (thought note that raster formats often have extra options, like how many dots per inch to use)."
+ "So far we talked about how to create and modify a single plot.\n",
+ "What if you have multiple plots you want to lay out in a certain way? You can do that. To place two plots next to each other, you can simply put them in a list and call `gggrid` on the list. Note that you first need to create the plots and save them as objects (in the following example they're called `p1` and `p2`).\n"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "d1492b5f",
+ "id": "a8081df4",
"metadata": {},
"outputs": [],
"source": [
- "(so.Plot(mpg, x=\"displ\", y=\"hwy\", color=\"class\").add(so.Dot()).save(\"output_chart.svg\"))"
+ "p1 = ggplot(mpg, aes(x=\"displ\", y=\"hwy\")) + geom_point() + labs(title=\"Plot 1\")\n",
+ "p2 = ggplot(mpg, aes(x=\"drv\", y=\"hwy\")) + geom_boxplot() + labs(title=\"Plot 2\")\n",
+ "gggrid([p1, p2])"
]
},
{
"cell_type": "markdown",
- "id": "6ca1b42b",
+ "id": "b0773270",
"metadata": {},
"source": [
- "To double check this works, let's use the terminal. We'll try the command `ls`, which lists everything in directory, and `grep *.svg` to pull out any files that end in `.svg` from what is returned by `ls`. These are strung together as commands by a `|`. (Note that the leading exclamation mark below just tells the software that builds this book to use the terminal.)"
+ "## Saving plots to file\n",
+ "\n",
+ "There are lots of output options to choose from to save your file to. Remember that, for graphics, *vector formats* are generally better than *raster formats*. In practice, this means saving plots in svg or pdf formats over jpg or png file formats. The svg format works in a lot of contexts (including Microsoft Word) and is a good default. To choose between formats, just supply the file extension and the file type will change automatically, eg \"chart.svg\" for svg or \"chart.png\" for png (thought note that raster formats often have extra options, like how many dots per inch to use).\n",
+ "\n",
+ "Let's try this out using the figure we made in the previous exercise, `p1`. `path=\".\"` just drops the file in the current directory."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "8ffc45b8",
+ "id": "710a6a4f",
"metadata": {},
"outputs": [],
"source": [
- "!ls | grep *.svg"
+ "ggsave(p1, \"chart.svg\", path=\".\")"
]
},
{
"cell_type": "markdown",
- "id": "549e2576",
+ "id": "7781794a",
"metadata": {},
"source": [
- "Great! It looks like our file saved successfully."
+ "To double check this has worked, let's use the terminal. We'll try the command `ls`, which lists everything in directory, and `grep *.svg` to pull out any files that end in `.svg` from what is returned by `ls`. These are strung together as commands by a `|`. (Note that the leading exclamation mark below just tells the software that builds this book to use the terminal.)"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "e7cf90a9",
+ "id": "bc831b1b",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "!ls | grep *.svg"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "9cc10ab7",
"metadata": {
"tags": [
"remove-cell"
@@ -455,7 +1314,22 @@
"# remove-cell\n",
"import os\n",
"\n",
- "os.remove(\"output_chart.svg\")"
+ "os.remove(\"chart.svg\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "793f4a04",
+ "metadata": {},
+ "source": [
+ "## Summary\n",
+ "\n",
+ "In this chapter you've learned about adding plot labels such as title, subtitle, caption as well as modifying default axis labels, using annotation to add informational text to your plot or to highlight specific data points, customising the axis scales, and changing the theme of your plot.\n",
+ "You've also learned about combining multiple plots in a single graph using both simple and complex plot layouts.\n",
+ "\n",
+ "While you've so far learned about how to make many different types of plots and how to customise them using a variety of techniques, we've barely scratched the surface of what you can create with **lets-plot**.\n",
+ "\n",
+ "The best place to go for further information is the [**lets-plot** dcoumentation](https://lets-plot.org/)."
]
}
],
@@ -484,7 +1358,7 @@
"name": "python",
"nbconvert_exporter": "python",
"pygments_lexer": "ipython3",
- "version": "3.9.12"
+ "version": "3.10.12"
},
"toc-showtags": true
},
diff --git a/data-transform.ipynb b/data-transform.ipynb
index afa8864..2468c80 100644
--- a/data-transform.ipynb
+++ b/data-transform.ipynb
@@ -135,6 +135,16 @@
"We would like to work with the `\"time_hour\"` variable in the form of a datetime; fortunately, **pandas** makes it easy to perform that conversion on that specific column"
]
},
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "ffb275b0",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "flights[\"time_hour\"]"
+ ]
+ },
{
"cell_type": "code",
"execution_count": null,
@@ -142,7 +152,7 @@
"metadata": {},
"outputs": [],
"source": [
- "flights[\"time_hour\"] = pd.to_datetime(flights[\"time_hour\"], format=\"%Y-%m-%d %H:%M:%S\")"
+ "flights[\"time_hour\"] = pd.to_datetime(flights[\"time_hour\"], format=\"%Y-%m-%dT%H:%M:%SZ\")"
]
},
{
@@ -1199,7 +1209,7 @@
"name": "python",
"nbconvert_exporter": "python",
"pygments_lexer": "ipython3",
- "version": "3.9.12"
+ "version": "3.10.12"
},
"toc-showtags": true
},
diff --git a/data-visualise.ipynb b/data-visualise.ipynb
index 4dab122..f2d890d 100644
--- a/data-visualise.ipynb
+++ b/data-visualise.ipynb
@@ -12,31 +12,13 @@
"\n",
"> \"The simple graph has brought more information to the data analyst's mind than any other device.\" --- John Tukey\n",
"\n",
- "This chapter will teach you how to visualise your data using the **seaborn** package.\n",
+ "This chapter will teach you how to visualise your data using using **[letsplot](https://lets-plot.org/)**.\n",
"\n",
- "There are a plethora of other options (and packages) for data visualisation using code. There are broadly two categories of approach to using code to create data visualisations: imperative, where you build what you want, and declarative, where you say what you want. Choosing which to use involves a trade-off: imperative libraries offer you flexibility but at the cost of some verbosity; declarative libraries offer you a quick way to plot your data, but only if it’s in the right format to begin with, and customisation may be more difficult.\n",
+ "There are broadly two categories of approach to using code to create data visualisations: imperative, where you build what you want, and declarative, where you say what you want. Choosing which to use involves a trade-off: imperative libraries offer you flexibility but at the cost of some verbosity; declarative libraries offer you a quick way to plot your data, but only if it’s in the right format to begin with, and customisation to special chart types is more difficult. Python has many excellent plotting packages, including perhaps the most powerful imperative plotting package around, **matplotlib**.\n",
"\n",
- "**seaborn** is a declarative visualisation package, and these can be easier to get started with. But it's built on top of an imperative package, the incredibly powerful **matplotlib**, so you can always dig further and tweak details if you need to. However, in this chapter, we'll focus on using **seaborn** declaratively."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "id": "51a55374",
- "metadata": {
- "tags": [
- "remove-cell"
- ]
- },
- "outputs": [],
- "source": [
- "# remove cell\n",
- "import matplotlib_inline.backend_inline\n",
- "import matplotlib.pyplot as plt\n",
+ "However, we'll get further faster by learning one system and applying it in many places—and the beauty of declarative plotting is that it covers lots of standard charts simply and well. **letsplot** implements the so-called **grammar of graphics**, a coherent declarative system for describing and building graphs.\n",
"\n",
- "# Plot settings\n",
- "plt.style.use(\"https://github.com/aeturrell/python4DS/raw/main/plot_style.txt\")\n",
- "matplotlib_inline.backend_inline.set_matplotlib_formats(\"svg\")"
+ "We will start by creating a simple scatterplot and use that to introduce aesthetic mappings and geometric objects—the fundamental building blocks of **letsplot**. We will then walk you through visualising distributions of single variables as well as visualising relationships between two or more variables. We’ll finish off with saving your plots and troubleshooting tips. "
]
},
{
@@ -46,17 +28,19 @@
"source": [
"### Prerequisites\n",
"\n",
- "You will need to install the **seaborn** package for this chapter (`pip install seaborn`). Once you've done this, you'll need to import the **seaborn** library into your session using"
+ "You will need to install the **letsplot** package for this chapter. To do this, open up the command line of your computer, type in `pip install letsplot`, and hit enter."
]
},
{
- "cell_type": "code",
- "execution_count": null,
- "id": "ae4a818a",
+ "cell_type": "markdown",
+ "id": "792902c7",
"metadata": {},
- "outputs": [],
"source": [
- "import seaborn.objects as so"
+ "```{note}\n",
+ "The command line can be opened within Visual Studio Code and Codespaces by going to View -> Terminal.\n",
+ "```\n",
+ "\n",
+ "Note that you only need to install a package once in each Python environment."
]
},
{
@@ -64,805 +48,1056 @@
"id": "e0ad70c8",
"metadata": {},
"source": [
- "The second import brings in the plotting part of **seaborn**.\n",
- "\n",
- "## First Steps\n",
+ "We'll also need to have the **pandas** package installed—this package, which we'll be seeing a lot of, is for data. You can similarly install it by running `pip install pandas` on the command line.\n",
"\n",
- "Let's use our first graph to answer a question: Do cars with big engines use more fuel than cars with small engines? You probably already have an answer, but try to make your answer precise. What does the relationship between engine size and fuel efficiency look like? Is it positive? Negative? Linear? Non-linear?\n",
- "\n",
- "### The `mpg` data frame\n",
- "\n",
- "You can test your answer with the `mpg` data frame found in **seaborn** and obtained from the internet using the **pandas** package.\n",
- "\n",
- "A data frame is a rectangular collection of variables (in the columns) and observations (in the rows). `mpg` contains observations collected by the US Environmental Protection Agency on 38 car models."
+ "Finally, we'll also need some data (you can't science with data). We'll be using the Palmer penguins dataset. Unusually, this can also be installed as a package—normally you would load data from a file, but these data are so popular for tutorials they've found their way into an installable package. Run `pip install palmerpenguins` to get these data."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "8852373a",
+ "metadata": {},
+ "source": [
+ "Our next task is to load these into our Python session, either in a Python notebook cell within a Jupyter Notebook, by writing it in a script that we then send to the interactive window, or by typing it directly into the interactive window and hitting shift and enter. Here's the code:"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "0cf986aa",
+ "id": "a86fb211",
"metadata": {},
"outputs": [],
"source": [
"import pandas as pd\n",
+ "from palmerpenguins import load_penguins\n",
+ "from lets_plot import *\n",
"\n",
- "mpg = pd.read_csv(\n",
- " \"https://vincentarelbundock.github.io/Rdatasets/csv/ggplot2/mpg.csv\", index_col=0\n",
- ")\n",
- "mpg"
+ "LetsPlot.setup_html()"
]
},
{
"cell_type": "markdown",
- "id": "cc310b4f",
+ "id": "4443f4dd",
"metadata": {},
"source": [
- "Among the variables in `mpg` are:\n",
- "\n",
- "1. `displ`, a car's engine size, in litres.\n",
+ "These lines import parts of the **pandas** and **palmerpenguins** packages, then import all (`*`) of the functions of the **letsplot** package. The final line allows charts to display in HTML."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "4bc87ab8",
+ "metadata": {},
+ "source": [
+ "## First Steps\n",
"\n",
- "2. `hwy`, a car's fuel efficiency on the highway, in miles per gallon (mpg). A car with a low fuel efficiency consumes more fuel than a car with a high fuel efficiency when they travel the same distance."
+ "Do penguins with longer flippers weigh more or less than penguins with shorter flippers? You probably already have an answer, but try to make your answer precise. What does the relationship between flipper length and body mass look like? Is it positive? Negative? Linear? Nonlinear? Does the relationship vary by the species of the penguin? How about by the island where the penguin lives? Let’s create visualisations that we can use to answer these questions."
]
},
{
"cell_type": "markdown",
- "id": "339966d7",
+ "id": "e4eb9c4f",
"metadata": {},
"source": [
- "### Creating a Plot\n",
+ "### The `penguins` data frame\n",
+ "\n",
+ "You can test your answers to those questions with the penguins data frame found in palmerpenguins (a.k.a. `from palmerpenguins import load_penguins`). A data frame is a rectangular collection of variables (in the columns) and observations (in the rows). `penguins` contains 344 observations collected and made available by Dr. Kristen Gorman and the Palmer Station, Antarctica LTER.{cite:p}`horst2020palmerpenguins`.\n",
+ "\n",
+ "To make the discussion easier, let's define some terms:\n",
+ "\n",
+ "- A **variable** is a quantity, quality, or property that you can measure.\n",
+ "\n",
+ "- A **value** is the state of a variable when you measure it.\n",
+ " The value of a variable may change from measurement to measurement.\n",
+ "\n",
+ "- An **observation** is a set of measurements made under similar conditions (you usually make all of the measurements in an observation at the same time and on the same object).\n",
+ " An observation will contain several values, each associated with a different variable.\n",
+ " We'll sometimes refer to an observation as a data point.\n",
+ "\n",
+ "- **Tabular data** is a set of values, each associated with a variable and an observation.\n",
+ " Tabular data is *tidy* if each value is placed in its own \"cell\", each variable in its own column, and each observation in its own row.\n",
+ "\n",
+ "In this context, a variable refers to an attribute of all the penguins, and an observation refers to all the attributes of a single penguin.\n",
"\n",
- "To plot `mpg`, run this code to put `displ` on the x-axis and `hwy` on the y-axis:"
+ "Type the name of the data frame in the interactive window and Python will print a preview of its contents.\n",
+ "Note that it says `shape` on top of this preview: that's the shape of your data (344 rows, 8 columns)."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "1b12b0ca",
+ "id": "0cf986aa",
"metadata": {},
"outputs": [],
"source": [
- "so.Plot(mpg, x=\"displ\", y=\"hwy\").add(so.Dot())"
+ "penguins = load_penguins()\n",
+ "penguins"
]
},
{
"cell_type": "markdown",
- "id": "7272e621",
+ "id": "cc310b4f",
"metadata": {},
"source": [
- "The plot shows a negative relationship between engine size (`displacement`) and fuel efficiency (`mpg`). In other words, cars with smaller engine sizes have higher fuel efficiency and, in general, as engine size increases, fuel efficiency decreases. Does this confirm or refute your hypothesis about fuel efficiency and engine size?\n",
- "\n",
- "With **seaborn**, you begin a plot with the function `so.Plot()`. **seaborn** creates a coordinate system that you can add layers to. The first argument of `so.Plot()` is the dataset to use in the graph. So `so.Plot(mpg)` creates an empty graph, but it's not very interesting so I'm not going to show it here.\n",
- "\n",
- "You complete your graph by adding one or more layers to the plot. The function `.add(so.Dot())` adds a layer of points to your plot, creating a scatterplot. You can choose between telling `so.Plot` what the x and y axis variables are or passing it directly to `.add`.\n",
- "\n",
- "**seaborn** comes with many functions that each add a different type of layer to a plot. You'll learn a whole bunch of them throughout this chapter."
+ "For an alternative view, where you can see the first few observations of each variable, use `penguins.head()`."
]
},
{
- "cell_type": "markdown",
- "id": "c5e295b2",
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "23c75ba7",
"metadata": {},
+ "outputs": [],
"source": [
- "### A graphing template\n",
- "\n",
- "Let's turn this code into a reusable template for making graphs with **seaborn**. To make a graph, replace the bracketed sections in the code below with a dataset, a geom function, or a collection of mappings.\n",
- "\n",
- "```python\n",
- "so.Plot(, x=, y=).add(so.)\n",
- "```\n",
- "\n",
- "The rest of this chapter will show you how to complete and extend this template to make different types of graphs."
+ "penguins.head()"
]
},
{
"cell_type": "markdown",
- "id": "351b59e2",
+ "id": "c3eb1881",
"metadata": {},
"source": [
- "### Exercises\n",
+ "Among the variables in `penguins` are:\n",
"\n",
- "1. Run `so.Plot(mpg)`.\n",
- " What do you see?\n",
+ "1. `species`: a penguin's species (Adelie, Chinstrap, or Gentoo).\n",
"\n",
- "2. How many rows are in `mpg` (the data frame)?\n",
- " How many columns?\n",
+ "2. `flipper_length_mm`: length of a penguin's flipper, in millimeters.\n",
"\n",
- "3. Make a scatterplot of `mpg` vs `cylinders`.\n",
+ "3. `body_mass_g`: body mass of a penguin, in grams.\n",
"\n",
- "4. What happens if you make a scatterplot of `class` vs `drv`? Why is the plot not useful?"
+ "To learn more about `penguins`, open the help page of its data-loading function by running `help(load_penguins)`.\n"
]
},
{
"cell_type": "markdown",
- "id": "e5867e3f",
+ "id": "caf04bde",
"metadata": {},
"source": [
- "## Aesthetic mappings\n",
+ "### Ultimate Goal\n",
"\n",
- "> \"The greatest value of a picture is when it forces us to notice what we never expected to see.\" --- John Tukey\n",
- "\n",
- "In the plot below, one group of points (highlighted in red) seems to fall outside of the linear trend. These cars have a higher mileage than you might expect. How can you explain these cars?\n"
+ "Our ultimate goal in this chapter is to recreate the following visualisation displaying the relationship between flipper lengths and body masses of these penguins, taking into consideration the species of the penguin."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "11877e4c",
+ "id": "574fe39f",
"metadata": {
"tags": [
- "remove-input"
+ "remove-cell"
]
},
"outputs": [],
"source": [
- "# remove input\n",
- "so.Plot(mpg, x=\"displ\", y=\"hwy\").add(so.Dot()).add(\n",
- " so.Dot(color=\"red\", pointsize=5), data=mpg.query(\"displ > 5 and hwy > 20\")\n",
+ "(\n",
+ " ggplot(penguins, aes(x=\"flipper_length_mm\", y=\"body_mass_g\"))\n",
+ " + geom_point(aes(color=\"species\", shape=\"species\"))\n",
+ " + geom_smooth(method=\"lm\")\n",
+ " + labs(\n",
+ " title=\"Body mass and flipper length\",\n",
+ " subtitle=\"Dimensions for Adelie, Chinstrap, and Gentoo Penguins\",\n",
+ " x=\"Flipper length (mm)\",\n",
+ " y=\"Body mass (g)\",\n",
+ " color=\"Species\",\n",
+ " shape=\"Species\",\n",
+ " )\n",
")"
]
},
{
"cell_type": "markdown",
- "id": "801606f0",
+ "id": "339966d7",
"metadata": {},
"source": [
- "Let's hypothesize that the cars are hybrids. One way to test this hypothesis is to look at the `class` value for each car.\n",
- "The `class` variable of the `mpg` dataset classifies cars into groups such as compact, midsize, and SUV. If the outlying points are hybrids, they should be classified as compact cars or, perhaps, subcompact cars (keep in mind that this data was collected before hybrid trucks and SUVs became popular).\n",
+ "### Creating a Plot\n",
+ "\n",
+ "Let's recreate this plot step-by-step.\n",
+ "\n",
+ "With **letsplot**, you begin a plot with the function `ggplot()`, defining a plot object that you then add **layers** to.\n",
+ "\n",
+ "The first argument of `ggplot()` is the dataset to use in the graph and so `ggplot(data = penguins)` creates an empty graph that is primed to display the `penguins` data, but since we haven't told it how to visualise it yet, for now it's empty. Because it's empty, running this alone would raise an error message: it's an empty canvas that you'll paint the remaining layers of your plot onto.\n",
+ "\n",
+ "```python\n",
+ "ggplot(data = penguins)\n",
+ "```\n",
+ "\n",
+ "Next, we need to tell `ggplot()` how the information from our data will be visually represented.\n",
+ "\n",
+ "The `mapping` argument of the `ggplot()` function defines how variables in your dataset are mapped to visual properties (**aesthetics**) of your plot.\n",
+ "The `mapping` argument is always defined in the `aes()` function, and the `x` and `y` arguments of `aes()` specify which variables to map to the x and y axes.\n",
+ "For now, we will only map flipper length to the `x` aesthetic and body mass to the `y` aesthetic. **letsplot** looks for the mapped variables in the `data` argument, in this case, `penguins`.\n",
+ "\n",
+ "Again, we haven't actually specified anything to plot, so running\n",
+ "\n",
+ "```python\n",
+ "ggplot(\n",
+ " data = penguins,\n",
+ " mapping = aes(x = \"flipper_length_mm\", y = \"body_mass_g\")\n",
+ ")\n",
+ "```\n",
+ "\n",
+ "would raise an error. This is because we have not yet articulated, in our code, how to represent the observations from our data frame on our plot.\n",
+ "\n",
+ "To do so, we need to define a **geom**: the geometrical object that a plot uses to represent data.\n",
+ "These geometric objects are made available in **letsplot** with functions that start with `geom_`.\n",
"\n",
- "You can add a third variable, like `class`, to a two dimensional scatterplot by mapping it to another dimension of the plot. These could be things like the size, the shape, or the colour of your points.\n",
+ "People often describe plots by the type of geom that the plot uses.\n",
+ "For example, bar charts use bar geoms (`geom_bar()`), line charts use line geoms (`geom_line()`), boxplots use boxplot geoms (`geom_boxplot()`), scatterplots use point geoms (`geom_point()`), and so on.\n",
"\n",
- "For example, you can map the colours of your points to the `class` variable to reveal the class of each car."
+ "The function `geom_point()` adds a layer of points to your plot, which creates a scatterplot.\n",
+ "**letsplot** comes with many geom functions that each adds a different type of layer to a plot."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "d3bd2335",
+ "id": "15c3848b",
"metadata": {},
"outputs": [],
"source": [
- "so.Plot(mpg, x=\"displ\", y=\"hwy\", color=\"class\").add(so.Dot())"
+ "(\n",
+ " ggplot(data=penguins, mapping=aes(x=\"flipper_length_mm\", y=\"body_mass_g\"))\n",
+ " + geom_point()\n",
+ ")"
]
},
{
"cell_type": "markdown",
- "id": "d4db1154",
+ "id": "c8d8bfff",
"metadata": {},
"source": [
- "To map another dimension in the plot to a variable, assign that dimension to the variable, for example `color=\"class\"` within `so.Plot` or within `.add`. **seaborn** will automatically assign a unique level of the dimension (here a unique colour) to each unique value of the variable, a process known as scaling. **seaborn** will also add a legend that explains which levels correspond to which values."
+ "Now we have something that looks like what we might think of as a \"scatterplot\".\n",
+ "It doesn't yet match our \"ultimate goal\" plot, but using this plot we can start answering the question that motivated our exploration: \"What does the relationship between flipper length and body mass look like?\" The relationship appears to be positive (as flipper length increases, so does body mass), fairly linear (the points are clustered around a line instead of a curve), and moderately strong (there isn't too much scatter around such a line).\n",
+ "Penguins with longer flippers are generally larger in terms of their body mass.\n",
+ "\n",
+ "It's a good point to flag that although we have plotted everything in the `penguins` data frame, there were a couple of rows with undefined values—and of course these cannot be plotted."
]
},
{
"cell_type": "markdown",
- "id": "9c4b3bb7",
+ "id": "40818ed6",
"metadata": {},
"source": [
- "The colours reveal that many of the unusual points (with engine size greater than 5 litres and highway fuel efficiency greater than 20 miles per gallon) are two-seater cars. These cars don't seem like hybrids, and are, in fact, sports cars! Sports cars have large engines like SUVs and pickup trucks, but small bodies like midsize and compact cars, which improves their gas mileage. In hindsight, these cars were unlikely to be hybrids since they have large engines.\n",
+ "### Adding aesthetics and layers\n",
+ "\n",
+ "Scatterplots are useful for displaying the relationship between two numerical variables, but it's always a good idea to be skeptical of any apparent relationship between two variables and ask if there may be other variables that explain or change the nature of this apparent relationship. For example, does the relationship between flipper length and body mass differ by species?\n",
+ "\n",
+ "Let's incorporate species into our plot and see if this reveals any additional insights into the apparent relationship between these variables.\n",
+ "We will do this by representing species with different colored points.\n",
"\n",
+ "To achieve this, will we need to modify the aesthetic or the geom?\n",
+ "If you guessed \"in the aesthetic mapping, inside of `aes()`\", you're already getting the hang of creating data visualisations with **letsplot**!\n",
+ "And if not, don't worry.\n",
"\n",
- "In the above example, we mapped `class` to colour, but we could have mapped `class` to the size of points in the same way. In this case, the exact size of each point would reveal its class affiliation. Big warning here though: mapping an unordered variable (`class`) to an ordered variable (`size`) is generally not a good idea."
+ "Throughout the book you will make many more plots and have many more opportunities to check your intuition as you make them."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "8ae59e98",
+ "id": "6b0e1c38",
"metadata": {},
"outputs": [],
"source": [
- "so.Plot(mpg, x=\"displ\", y=\"hwy\", pointsize=\"class\").add(so.Dot())"
+ "(\n",
+ " ggplot(\n",
+ " data=penguins,\n",
+ " mapping=aes(x=\"flipper_length_mm\", y=\"body_mass_g\", color=\"species\"),\n",
+ " )\n",
+ " + geom_point()\n",
+ ")"
]
},
{
"cell_type": "markdown",
- "id": "c1c17016",
+ "id": "7272e621",
"metadata": {},
"source": [
- "Similarly, we could have mapped `class` to *alpha* level, which controls the transparency of the points, or to the *marker* variable, which controls the shape of the points."
+ "When a categorical variable is mapped to an aesthetic, **letsplot** will automatically assign a unique value of the aesthetic (here a unique color) to each unique level of the variable (each of the three species), a process known as **scaling**.\n",
+ "\n",
+ "**letsplot** will also add a legend that explains which values correspond to which levels.\n",
+ "\n",
+ "Now let's add one more layer: a smooth curve displaying the relationship between body mass and flipper length.\n",
+ "\n",
+ "Before you proceed, refer back to the code above, and think about how we can add this to our existing plot.\n",
+ "\n",
+ "Since this is a new geometric object representing our data, we will add a new geom as a layer on top of our point geom: `geom_smooth()`.\n",
+ "\n",
+ "And we will specify that we want to draw the line of best fit based on a `l`inear `m`odel with `method = \"lm\"`."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "dcfa90ee",
+ "id": "943efd36",
"metadata": {},
"outputs": [],
"source": [
- "so.Plot(mpg, x=\"displ\", y=\"hwy\", alpha=\"class\").add(so.Dot())"
+ "(\n",
+ " ggplot(\n",
+ " data=penguins,\n",
+ " mapping=aes(x=\"flipper_length_mm\", y=\"body_mass_g\", color=\"species\"),\n",
+ " )\n",
+ " + geom_point()\n",
+ " + geom_smooth(method=\"lm\")\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "3c1cd3c7",
+ "metadata": {},
+ "source": [
+ "We have successfully added lines, but this plot doesn't look like the plot from earlier as that only had one line for the entire dataset as opposed to separate lines for each of the penguin species.\n",
+ "\n",
+ "When aesthetic mappings are defined in `ggplot()`, at the *global* level, they're passed down to each of the subsequent geom layers of the plot.\n",
+ "\n",
+ "However, each geom function in **letplot** can also take a `mapping` argument, which allows for aesthetic mappings at the *local* level that are added to those inherited from the global level.\n",
+ "\n",
+ "Since we want points to be colored based on species but don't want the lines to be separated out for them, we should specify `color = species` for `geom_point()` only: therefore we take it out of the global `aes` and just add it to `geom_point()`.\n"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "824214a5",
+ "id": "9e12b3bf",
"metadata": {},
"outputs": [],
"source": [
- "so.Plot(mpg, x=\"displ\", y=\"hwy\", marker=\"class\").add(so.Dot())"
+ "(\n",
+ " ggplot(data=penguins, mapping=aes(x=\"flipper_length_mm\", y=\"body_mass_g\"))\n",
+ " + geom_point(mapping=aes(color=\"species\"))\n",
+ " + geom_smooth(method=\"lm\")\n",
+ ")"
]
},
{
"cell_type": "markdown",
- "id": "a637a55b",
+ "id": "928898ef",
"metadata": {},
"source": [
- "Once you map variables to dimensions, **seaborn** takes care of the rest. It selects a reasonable scale to use with the dimension, and it constructs a legend that explains the mapping between levels and values.\n",
+ "Voila! We have something that looks very much like our ultimate goal, though it's not yet perfect.\n",
+ "\n",
+ "We still need to use different shapes for each species of penguins and improve labels.\n",
"\n",
- "You can also *set* a dimension property in your plot directly. For example, we can make all of the points in our plot purple:"
+ "It's generally not a good idea to represent information using only colors on a plot, as people perceive colors differently due to color blindness or other color vision differences. Therefore, in addition to color, we can also map `species` to the `shape` aesthetic."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "b3d9fb32",
+ "id": "17d5803b",
"metadata": {},
"outputs": [],
"source": [
- "so.Plot(mpg, x=\"displ\", y=\"hwy\").add(so.Dot(color=\"purple\"))"
+ "(\n",
+ " ggplot(data=penguins, mapping=aes(x=\"flipper_length_mm\", y=\"body_mass_g\"))\n",
+ " + geom_point(mapping=aes(color=\"species\", shape=\"species\"))\n",
+ " + geom_smooth(method=\"lm\")\n",
+ ")"
]
},
{
"cell_type": "markdown",
- "id": "af32bb82",
+ "id": "3cfae7fc",
"metadata": {},
"source": [
- "Here, the colour doesn't convey information about a variable, but only changes the appearance of the plot.\n",
- "To set a dimension manually like this, put it within the specific layer it applies to (eg `.add(so.Scatter(color=\"purple\"))`) rather than in the part that maps variables to dimensions (eg not in `so.Plot(mpg, x=\"displ\", y=\"hwy\")`).\n",
- "\n",
- "When assigning values to dimensions, you'll need to pick values that makes sense, for example:\n",
+ "Note that the legend is automatically updated to reflect the different shapes of the points as well.\n",
"\n",
- "- The name of a colour as a string, eg `color=\"purple\"`\n",
- "- The size of a point in mm\n",
- "- The shape of a marker as a string, eg `marker=\"*\"` for a star"
+ "And finally, we can improve the labels of our plot using the `labs()` function in a new layer. Some of the arguments to `labs()` might be self explanatory: `title` adds a title and `subtitle` adds a subtitle to the plot. Other arguments match the aesthetic mappings, `x` is the x-axis label, `y` is the y-axis label, and `color` and `shape` define the label for the legend."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "cc31c7f4",
- "metadata": {
- "tags": [
- "remove-cell"
- ]
- },
+ "id": "b9b98ec4",
+ "metadata": {},
"outputs": [],
"source": [
- "# remove cell\n",
- "from matplotlib.lines import Line2D\n",
- "\n",
- "\n",
- "text_style = dict(\n",
- " horizontalalignment=\"right\",\n",
- " verticalalignment=\"center\",\n",
- " fontsize=12,\n",
- " fontfamily=\"monospace\",\n",
- ")\n",
- "marker_style = dict(\n",
- " linestyle=\":\",\n",
- " color=\"0.8\",\n",
- " markersize=10,\n",
- " markerfacecolor=\"tab:blue\",\n",
- " markeredgecolor=\"tab:blue\",\n",
- ")\n",
- "\n",
- "\n",
- "def format_axes(ax):\n",
- " ax.margins(0.2)\n",
- " ax.set_axis_off()\n",
- " ax.invert_yaxis()\n",
- "\n",
- "\n",
- "def split_list(a_list):\n",
- " i_half = len(a_list) // 2\n",
- " return a_list[:i_half], a_list[i_half:]\n",
- "\n",
- "\n",
- "fig, axs = plt.subplots(ncols=2)\n",
- "fig.suptitle(\"Un-filled markers\", fontsize=14)\n",
- "\n",
- "# Filter out filled markers and marker settings that do nothing.\n",
- "unfilled_markers = [\n",
- " m\n",
- " for m, func in Line2D.markers.items()\n",
- " if func != \"nothing\" and m not in Line2D.filled_markers\n",
- "]\n",
- "\n",
- "for ax, markers in zip(axs, split_list(unfilled_markers)):\n",
- " for y, marker in enumerate(markers):\n",
- " ax.text(-0.5, y, repr(marker), **text_style)\n",
- " ax.plot([y] * 3, marker=marker, **marker_style)\n",
- " format_axes(ax)\n",
- "\n",
- "plt.show()\n",
- "\n",
- "fig, axs = plt.subplots(ncols=2)\n",
- "fig.suptitle(\"Filled markers\", fontsize=14)\n",
- "for ax, markers in zip(axs, split_list(Line2D.filled_markers)):\n",
- " for y, marker in enumerate(markers):\n",
- " ax.text(-0.5, y, repr(marker), **text_style)\n",
- " ax.plot([y] * 3, marker=marker, **marker_style)\n",
- " format_axes(ax)\n",
- "\n",
- "plt.show()"
+ "(\n",
+ " ggplot(data=penguins, mapping=aes(x=\"flipper_length_mm\", y=\"body_mass_g\"))\n",
+ " + geom_point(aes(color=\"species\", shape=\"species\"))\n",
+ " + geom_smooth(method=\"lm\")\n",
+ " + labs(\n",
+ " title=\"Body mass and flipper length\",\n",
+ " subtitle=\"Dimensions for Adelie, Chinstrap, and Gentoo Penguins\",\n",
+ " x=\"Flipper length (mm)\",\n",
+ " y=\"Body mass (g)\",\n",
+ " color=\"Species\",\n",
+ " shape=\"Species\",\n",
+ " )\n",
+ ")"
]
},
{
"cell_type": "markdown",
- "id": "42b0c66e",
+ "id": "cdc33b33",
"metadata": {},
"source": [
- "You can find more information on markers in the [**matplotlib** documentation](https://matplotlib.org/stable/gallery/lines_bars_and_markers/marker_reference.html)"
+ "We finally have a plot that perfectly matches our \"ultimate goal\"!"
]
},
{
"cell_type": "markdown",
- "id": "fc7ac561",
+ "id": "81863a95",
"metadata": {},
"source": [
- "## Facets\n",
+ "### Exercises\n",
"\n",
- "One way to add additional variables to a plot is by mapping them to a dimension. Another way, which is particularly useful for categorical variables, is to split your plot into **facets**, subplots that each display one subset of the data.\n",
+ "1. How many rows are in `penguins`?\n",
+ " How many columns?\n",
"\n",
- "To facet your plot by a single variable, use `.facet()`; this should be a discrete variable."
+ "2. What does the `bill_depth_mm` variable in the `penguins` data frame describe?\n",
+ " Read the help for `?penguins` to find out.\n",
+ "\n",
+ "3. Make a scatterplot of `bill_depth_mm` vs. `bill_length_mm`.\n",
+ " That is, make a scatterplot with `bill_depth_mm` on the y-axis and `bill_length_mm` on the x-axis.\n",
+ " Describe the relationship between these two variables.\n",
+ "\n",
+ "4. What happens if you make a scatterplot of `species` vs. `bill_depth_mm`?\n",
+ " What might be a better choice of geom?\n",
+ "\n",
+ "5. Why does the following give an error and how would you fix it?\n",
+ "\n",
+ " ```python\n",
+ " (ggplot(data = penguins) + \n",
+ " geom_point())\n",
+ " ```\n",
+ "\n",
+ "6. Add the following caption to the plot you made in the previous exercise: \"Data come from the palmerpenguins package.\" Hint: Take a look at the documentation for `labs()`.\n",
+ "\n",
+ "7. Recreate the following visualisation.\n",
+ " What aesthetic should `bill_depth_mm` be mapped to?\n",
+ " And should it be mapped at the global level or at the geom level?"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "e1fc6ec1",
- "metadata": {},
+ "id": "7c76be4b",
+ "metadata": {
+ "tags": [
+ "remove-cell"
+ ]
+ },
"outputs": [],
"source": [
"(\n",
- " so.Plot(\n",
- " mpg,\n",
- " \"displ\",\n",
- " \"hwy\",\n",
- " )\n",
- " .facet(\"cyl\")\n",
- " .add(so.Dot())\n",
+ " ggplot(data=penguins, mapping=aes(x=\"flipper_length_mm\", y=\"body_mass_g\"))\n",
+ " + geom_point(aes(color=\"bill_depth_mm\"))\n",
+ " + geom_smooth()\n",
")"
]
},
{
"cell_type": "markdown",
- "id": "ed1d597e",
+ "id": "986fdc29",
"metadata": {},
"source": [
- "## Geometric objects\n",
"\n",
- "How are these two plots similar?"
+ "8. Run this code in your head and predict what the output will look like.\n",
+ " Then, run the code in Python and check your predictions.\n",
+ "\n",
+ " ```python\n",
+ "\n",
+ " (ggplot(\n",
+ " data = penguins,\n",
+ " mapping = aes(x = \"flipper_length_mm\", y = \"body_mass_g\", color = \"island\")\n",
+ " ) +\n",
+ " geom_point() +\n",
+ " geom_smooth(se = False)\n",
+ " )\n",
+ " ```\n",
+ "\n",
+ "9. Will these two graphs look different?\n",
+ " Why/why not?\n",
+ "\n",
+ " ```python\n",
+ "\n",
+ " (ggplot(\n",
+ " data = penguins,\n",
+ " mapping = aes(x = \"flipper_length_mm\", y = \"body_mass_g\")\n",
+ " ) +\n",
+ " geom_point() +\n",
+ " geom_smooth()\n",
+ " )\n",
+ " ```\n",
+ " ```python\n",
+ " (ggplot() +\n",
+ " geom_point(\n",
+ " data = penguins,\n",
+ " mapping = aes(x = \"flipper_length_mm\", y = \"body_mass_g\")\n",
+ " ) +\n",
+ " geom_smooth(\n",
+ " data = penguins,\n",
+ " mapping = aes(x = \"flipper_length_mm\", y = \"body_mass_g\")\n",
+ " )\n",
+ " )\n",
+ " ```"
]
},
{
- "cell_type": "code",
- "execution_count": null,
- "id": "c8fa9864",
+ "cell_type": "markdown",
+ "id": "10806a67",
"metadata": {},
- "outputs": [],
"source": [
- "so.Plot(mpg, x=\"displ\", y=\"hwy\").add(so.Dot())"
+ "## **letsplot** calls\n",
+ "\n",
+ "As we move on from these introductory sections, we'll transition to a more concise expression of **letsplot** code.\n",
+ "\n",
+ "So far we've been very explicit, which is helpful when you are learning:\n",
+ "\n",
+ "```python\n",
+ "(ggplot(\n",
+ " data = penguins,\n",
+ " mapping = aes(x = \"flipper_length_mm\", y = \"body_mass_g\")\n",
+ ") +\n",
+ " geom_point())\n",
+ "```"
]
},
{
- "cell_type": "code",
- "execution_count": null,
- "id": "693f73ea",
+ "cell_type": "markdown",
+ "id": "e4f403fb",
"metadata": {},
- "outputs": [],
"source": [
- "so.Plot(mpg, x=\"displ\", y=\"hwy\").add(so.Line(), so.Agg())"
+ "Typically, the first one or two arguments to a function are so important that you should know them by heart.\n",
+ "The first two arguments to `ggplot()` are `data` and `mapping`, in the remainder of the book, we won't supply those names—the way the function is written, Python knows to expect these variables because of their position. Not writing them in saves typing, and, by reducing the amount of extra text, makes it easier to see what's different between plots.\n",
+ "That's a really important programming concern that we'll come back to later.\n",
+ "\n",
+ "Rewriting the previous plot more concisely yields:\n",
+ "\n",
+ "```python\n",
+ "(\n",
+ " ggplot(penguins, aes(x = \"flipper_length_mm\", y = \"body_mass_g\")) + \n",
+ " geom_point()\n",
+ ")\n",
+ "```"
]
},
{
"cell_type": "markdown",
- "id": "c966f9e0",
+ "id": "8d219ea2",
"metadata": {},
"source": [
- "Both plots contain the same x variable, the same y variable, and both show the same data (to some extent). But the plots are not identical. Each plot uses a different visual object to represent the data. In **seaborn** language, these are represented by different *marks*: one is a scatter and the other a (mean) line (which introduces an aggregation).\n",
+ "## visualising distributions\n",
+ "\n",
+ "How you visualise the distribution of a variable depends on the type of variable: categorical or numerical.\n",
+ "\n",
+ "### A categorical variable\n",
"\n",
- "A mark is a geometrical object that shows where data occur in x, y, and any other dimension-space you care to use. For example, the plot below is a line plot but we've added a discrete dimension of colour so that—instead of a single aggregate line—we get one for each distinct value of `\"drv\"`. One line describes all of the points that have a `4` value, one line describes all of the points that have an `f` value, and one line describes all of the points that have an `r` value. Here, `4` stands for four-wheel drive, `f` for front-wheel drive, and `r` for rear-wheel drive."
+ "A variable is **categorical** if it can only take one of a small set of values.\n",
+ "To examine the distribution of a categorical variable, you can use a bar chart.\n",
+ "The height of the bars displays how many observations occurred with each `x` value.\n"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "3067686e",
+ "id": "21b45061",
"metadata": {},
"outputs": [],
"source": [
- "so.Plot(mpg, x=\"displ\", y=\"hwy\", color=\"drv\").add(so.Line(), so.Agg())"
+ "(ggplot(penguins, aes(x=\"species\")) + geom_bar())"
]
},
{
"cell_type": "markdown",
- "id": "84a07969",
+ "id": "699f42eb",
"metadata": {},
"source": [
- "You can achieve the same effect without distinguishing by colour too using `group` keyword. The `group` keyword uses a categorical variable to draw multiple objects; **seaborn** will draw a separate object for each unique value of the grouping variable.\n"
+ "You may have seen earlier that the *data type* of the `\"species\"` column is string. Ideally, we want it to be categorical, so that there's no confusion about the fact that we're dealing with a finite number of mutually exclusive groups here. Another advantage is that it allows plotting tools to realise what kind of data it is working with.\n",
+ "\n",
+ "We can transform the variable to a categorical variable using **pandas** like so:"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "af6a8a64",
+ "id": "4e046bb2",
"metadata": {},
"outputs": [],
"source": [
- "so.Plot(mpg, x=\"displ\", y=\"hwy\").add(so.Line(), so.Agg(), group=\"drv\")"
+ "penguins[\"species\"] = penguins[\"species\"].astype(\"category\")\n",
+ "penguins.head()"
]
},
{
"cell_type": "markdown",
- "id": "4aadf721",
+ "id": "06d834a5",
"metadata": {},
"source": [
- "**seaborn** will allow you to add multiple layers to the base plot. In the below, we show both the points (using `.add(so.Dot())`) and an aggregate line per value of `\"drv\"`. Because we passed colour into `.Plot` both of these layers are distinguished by different colours."
+ "You will learn more about categorical variables later in the book."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "f9ca3124",
+ "metadata": {},
+ "source": [
+ "\n",
+ "### A numerical variable\n",
+ "\n",
+ "A variable is **numerical** (or quantitative) if it can take on a wide range of numerical values, and it is sensible to add, subtract, or take averages with those values. Numerical variables can be continuous or discrete.\n",
+ "\n",
+ "One commonly used visualisation for distributions of continuous variables is a histogram."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "571fedc3",
+ "id": "93675336",
"metadata": {},
"outputs": [],
"source": [
- "so.Plot(mpg, x=\"displ\", y=\"hwy\", color=\"drv\").add(so.Line(), so.Agg()).add(so.Dot())"
+ "(ggplot(penguins, aes(x=\"body_mass_g\")) + geom_histogram(binwidth=200))"
]
},
{
"cell_type": "markdown",
- "id": "f6caa33b",
+ "id": "cdac30fd",
"metadata": {},
"source": [
- "If you map variables to the dimensions in `.Plot`, **seaborn** will use them for all subsequent layers. But if you specify a different approach for a specific layer, you will get info just for that:"
+ "A histogram divides the x-axis into equally spaced bins and then uses the height of a bar to display the number of observations that fall in each bin.\n",
+ "In the graph above, the tallest bar shows that 39 observations have a `body_mass_g` value between 3,500 and 3,700 grams, which are the left and right edges of the bar.\n",
+ "\n",
+ "You can set the width of the intervals in a histogram with the binwidth argument, which is measured in the units of the `x` variable.\n",
+ "You should always explore a variety of binwidths when working with histograms, as different binwidths can reveal different patterns.\n",
+ "In the plots below a binwidth of 20 is too narrow, resulting in too many bars, making it difficult to determine the shape of the distribution.\n",
+ "Similarly, a binwidth of 2,000 is too high, resulting in all data being binned into only three bars, and also making it difficult to determine the shape of the distribution.\n",
+ "A binwidth of 200 provides a sensible balance, but you should always look at your data a few different ways, especially with histograms as they can be misleading."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "a5f1980f",
+ "metadata": {},
+ "source": [
+ "An alternative visualisation for distributions of numerical variables is a density plot.\n",
+ "A density plot is a smoothed-out version of a histogram and a practical alternative, particularly for continuous data that comes from an underlying smooth distribution.\n",
+ "We won't go into how `geom_density()` estimates the density (you can read more about that in the function documentation), but let's explain how the density curve is drawn with an analogy.\n",
+ "Imagine a histogram made out of wooden blocks.\n",
+ "Then, imagine that you drop a cooked spaghetti string over it.\n",
+ "The shape the spaghetti will take draped over blocks can be thought of as the shape of the density curve.\n",
+ "It shows fewer details than a histogram but can make it easier to quickly glean the shape of the distribution, particularly with respect to modes and skewness.\n"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "e1c59f65",
+ "id": "6a58021f",
"metadata": {},
"outputs": [],
"source": [
- "so.Plot(mpg, x=\"displ\", y=\"hwy\").add(so.Line(), so.Agg()).add(so.Dot(), color=\"class\")"
+ "(ggplot(penguins, aes(x=\"body_mass_g\")) + geom_density())"
]
},
{
"cell_type": "markdown",
- "id": "68e8a713",
+ "id": "f0086086",
"metadata": {},
"source": [
- "Each layer can have its own cut of the data too. Here, our line displays just a subset of the `mpg` dataset, the subcompact cars. We get this by explicitly adding a `data=` keyword argument to the same `.add` command as a line. The scatter plot has all points, the line just those for subcompact cars as specified by the filter we applied to the **pandas** data frame (try running `mpg.loc[mpg[\"class\"] == \"subcompact\"]` to see the data that make up the line)."
+ "### Exercises\n",
+ "\n",
+ "1. Make a bar plot of `\"species\"` of `penguins`, where you assign `\"species\"` to the `y` aesthetic.\n",
+ " How is this plot different?\n",
+ "\n",
+ "2. How are the following two plots different?\n",
+ " Which aesthetic, `color` or `fill`, is more useful for changing the color of bars?\n",
+ "\n",
+ " ```Python\n",
+ "\n",
+ " (ggplot(penguins, aes(x = species)) +\n",
+ " geom_bar(color = \"red\"))\n",
+ "\n",
+ " (ggplot(penguins, aes(x = species)) +\n",
+ " geom_bar(fill = \"red\"))\n",
+ " ```\n",
+ "\n",
+ "3. What does the `bins` argument in `geom_histogram()` do?"
]
},
{
- "cell_type": "code",
- "execution_count": null,
- "id": "8606b4fb",
+ "cell_type": "markdown",
+ "id": "015b31a0",
"metadata": {},
- "outputs": [],
"source": [
- "(\n",
- " so.Plot(mpg, x=\"displ\", y=\"hwy\")\n",
- " .add(so.Dot())\n",
- " .add(so.Line(color=\"blue\"), so.Agg(), data=mpg.loc[mpg[\"class\"] == \"subcompact\"])\n",
- ")"
+ "## Visualising Relationships\n",
+ "\n",
+ "To visualise a relationship we need to have at least two variables mapped to aesthetics of a plot—though you should remember that correlation is not causation, and causation is not correlation!\n",
+ "\n",
+ "In the following sections you will learn about commonly used plots for visualising relationships between two or more variables and the geoms used for creating them."
]
},
{
"cell_type": "markdown",
- "id": "a3e3b57a",
+ "id": "85458170",
"metadata": {},
"source": [
- "## Statistical Transformations\n",
+ "### A numerical and a categorical variable\n",
+ "\n",
+ "To visualise the relationship between a numerical and a categorical variable we can use side-by-side box plots.\n",
+ "\n",
+ "A **boxplot** is a type of visual shorthand for measures of position within a distribution (percentiles).\n",
+ "\n",
+ "It is also useful for identifying potential outliers. Each boxplot consists of:\n",
"\n",
- "We've already seen `so.Agg()` for aggregating multiple points into a single, mean line. Now let's take a look at another statistical transform: the bar chart. We'll use the diamonds dataset:"
+ "- A box that indicates the range of the middle half of the data, a distance known as the interquartile range (IQR), stretching from the 25th percentile of the distribution to the 75th percentile.\n",
+ " In the middle of the box is a line that displays the median, i.e. 50th percentile, of the distribution.\n",
+ " These three lines give you a sense of the spread of the distribution and whether or not the distribution is symmetric about the median or skewed to one side.\n",
+ "\n",
+ "- Visual points that display observations that fall more than 1.5 times the IQR from either edge of the box.\n",
+ " These outlying points are unusual so are plotted individually.\n",
+ "\n",
+ "- A line (or whisker) that extends from each end of the box and goes to the farthest non-outlier point in the distribution.\n",
+ "\n",
+ "\n",
+ "Let's take a look at the distribution of body mass by species using `geom_boxplot()`:"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "e0e025d8",
+ "id": "a636947a",
"metadata": {},
"outputs": [],
"source": [
- "import seaborn as sns\n",
- "\n",
- "diamonds = sns.load_dataset(\"diamonds\")\n",
- "diamonds.head()"
+ "(ggplot(penguins, aes(x=\"species\", y=\"body_mass_g\")) + geom_boxplot())"
]
},
{
"cell_type": "markdown",
- "id": "947113bd",
+ "id": "97b24caa",
"metadata": {},
"source": [
- "Let's now create a bar chart of counts, aka a histogram, of the numbers of diamonds of different cuts. This only requires one dimension, `\"cut\"`, and then an instruction to use `so.Hist()` alongside `so.Bar()` in the (single) layer on top of the plot."
+ "Alternatively, we can make probability density plots with `geom_density()`."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "6d063023",
+ "id": "9b85a2df",
"metadata": {},
"outputs": [],
"source": [
- "(so.Plot(diamonds, \"cut\").add(so.Bar(), so.Hist()))"
+ "(ggplot(penguins, aes(x=\"body_mass_g\", color=\"species\")) + geom_density(size=2))"
]
},
{
"cell_type": "markdown",
- "id": "0d91f0f7",
+ "id": "e0c10ae4",
"metadata": {},
"source": [
- "On the x-axis, the chart displays cut, a variable from diamonds. On the y-axis, it displays count, but count is not a variable in diamonds! Where does count come from? Many graphs, like scatterplots, plot the raw values of your dataset. Other graphs, like bar charts, calculate new values to plot:\n",
+ "We've also customized the thickness of the lines using the `size` argument in order to make them stand out a bit more against the background.\n",
"\n",
- "- bar charts, histograms, and frequency polygons bin your data and then plot bin counts, the number of points that fall in each bin\n",
- "\n",
- "- aggregations fit a mean line to your data \n",
- "\n",
- "- boxplots compute a summary of the distribution and display it as a box\n",
- "\n",
- "The algorithm used to calculate new values for a graph is called a Stat, short for statistical transformation."
+ "Additionally, we can map `species` to both `color` and `fill` aesthetics and use the `alpha` aesthetic to add transparency to the filled density curves.\n",
+ "This aesthetic takes values between 0 (completely transparent) and 1 (completely opaque).\n",
+ "In the following plot it's *set* to 0.5."
]
},
{
- "cell_type": "markdown",
- "id": "bdf9ec82",
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "353189e5",
"metadata": {},
+ "outputs": [],
"source": [
- "## Adding More Information to Plots\n",
- "\n",
- "\n",
- "There’s one more piece of magic associated with bar charts. You can colour a bar chart using the `color=` keyword argument within the `.add` layer"
+ "(\n",
+ " ggplot(penguins, aes(x=\"body_mass_g\", color=\"species\", fill=\"species\"))\n",
+ " + geom_density(alpha=0.5)\n",
+ ")"
]
},
{
- "cell_type": "code",
- "execution_count": null,
- "id": "04a0fb17",
+ "cell_type": "markdown",
+ "id": "0a2c7d59",
"metadata": {},
- "outputs": [],
"source": [
- "(so.Plot(diamonds, \"cut\").add(so.Bar(), so.Hist(), color=\"cut\"))"
+ "Note the terminology we have used here:\n",
+ "\n",
+ "- We *map* variables to aesthetics if we want the visual attribute represented by that aesthetic to vary based on the values of that variable.\n",
+ "- Otherwise, we *set* the value of an aesthetic.\n"
]
},
{
"cell_type": "markdown",
- "id": "2953116b",
+ "id": "63de3309",
"metadata": {},
"source": [
- "But you can also choose another variable and thereby add extra info to your chart, for example here by adding information on clarity:"
+ "### Two categorical variables\n",
+ "\n",
+ "We can use stacked bar plots to visualise the relationship between two categorical variables.\n",
+ "\n",
+ "For example, the following two stacked bar plots both display the relationship between `island` and `species`, or specifically, visualising the distribution of `species` within each island.\n",
+ "\n",
+ "The first plot shows the frequencies of each species of penguins on each island.\n",
+ "The plot of frequencies show that there are equal numbers of Adelies on each island.\n",
+ "\n",
+ "But we don't have a good sense of the percentage balance within each island."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "f1bdfcbb",
+ "id": "e091e211",
"metadata": {},
"outputs": [],
"source": [
- "(so.Plot(diamonds, \"cut\").add(so.Bar(), so.Hist(), color=\"clarity\"))"
+ "(ggplot(penguins, aes(x=\"island\", fill=\"species\")) + geom_bar())"
]
},
{
"cell_type": "markdown",
- "id": "d4abe5f1",
+ "id": "8e34c211",
"metadata": {},
"source": [
- "### Overplotting\n",
+ "The second plot is a relative frequency plot, created by setting `position = \"fill\"` in the geom is more useful for comparing species distributions across islands since it's not affected by the unequal numbers of penguins across the islands.\n",
"\n",
- "**Seaborn** functions have parameters that allow adjustments for overplotting, ie putting multiple dimensions next to each other on the same chart. These include `dodge` in several categorical functions, `jitter` in several functions based on scatterplots, and the `multiple=` parameter in distribution functions. These adjustments are abstracted away from the particular visual representation into the concept of a 'move':"
+ "Using this plot we can see that Gentoo penguins all live on Biscoe island and make up roughly 75% of the penguins on that island, Chinstrap all live on Dream island and make up roughly 50% of the penguins on that island, and Adelie live on all three islands and make up all of the penguins on Torgersen.\n"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "912c8045",
+ "id": "7df8fb7a",
"metadata": {},
"outputs": [],
"source": [
- "(so.Plot(diamonds, \"cut\", color=\"clarity\").add(so.Bar(), so.Hist(), so.Dodge()))"
+ "(ggplot(penguins, aes(x=\"island\", fill=\"species\")) + geom_bar(position=\"fill\"))"
]
},
{
"cell_type": "markdown",
- "id": "e68dc4e1",
- "metadata": {},
- "source": [
- "This can also accept parameters to separate out the information in a particular way"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "id": "999346aa",
+ "id": "cc83c3db",
"metadata": {},
- "outputs": [],
"source": [
- "(\n",
- " so.Plot(diamonds, \"cut\", color=\"clarity\").add(\n",
- " so.Bar(), so.Hist(), so.Dodge(empty=\"fill\", gap=0.5)\n",
- " )\n",
- ")"
+ "In creating these bar charts, we map the variable that will be separated into bars to the `x` aesthetic, and the variable that will change the colors inside the bars to the `fill` aesthetic."
]
},
{
"cell_type": "markdown",
- "id": "63edd20b",
+ "id": "f77e5c39",
"metadata": {},
"source": [
- "There's another type of adjustment that's not useful for bar charts, but can be very useful for scatterplots. Recall our first scatterplot. Did you notice that the plot displays only 126 points, even though there are 234 observations in the dataset? And that some points appear darker than others?"
+ "### Two numerical variables\n",
+ "\n",
+ "So far you've learned about scatterplots (created with `geom_point()`) and smooth curves (created with `geom_smooth()`) for visualising the relationship between two numerical variables.\n",
+ "A scatterplot is probably the most commonly used plot for visualising the relationship between two numerical variables.\n"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "1d06805f",
+ "id": "5066527d",
"metadata": {},
"outputs": [],
"source": [
- "so.Plot(mpg, x=\"displ\", y=\"hwy\").add(so.Dot())"
+ "(ggplot(penguins, aes(x=\"flipper_length_mm\", y=\"body_mass_g\")) + geom_point())"
]
},
{
"cell_type": "markdown",
- "id": "b519751a",
+ "id": "427f22c9",
"metadata": {},
"source": [
+ "### Three or more variables\n",
"\n",
- "The underlying values of `hwy` and `displ` are rounded so the points appear on a grid and many points overlap each other. This problem is known as **overplotting**. This arrangement makes it difficult to see the distribution of the data. Because scatterplot points are, by default, plotted with some transparency you can get a sense of which parts of the grid have multiple points on them, but you may wish to use a different technique.\n",
+ "As we saw already, we can incorporate more variables into a plot by mapping them to additional aesthetics.\n",
"\n",
- "Another way to show the overlap is to use the \"jitter\" option. Passing the argument `so.Jitter()` adds a small amount of random noise to each point. Depending on the numerical option you use, this spreads the points out because no two points are likely to receive the same amount of random noise.\n"
+ "For example, in the following scatterplot the colors of points represent species and the shapes of points represent islands.\n"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "d9f943f2",
+ "id": "8ca23d34",
"metadata": {},
"outputs": [],
"source": [
- "so.Plot(mpg, x=\"displ\", y=\"hwy\").add(so.Dot(), so.Jitter(1))"
- ]
- },
- {
- "cell_type": "markdown",
- "id": "7db1dfec",
- "metadata": {},
- "source": [
- "Adding randomness seems like a strange way to improve your plot, but while it makes your graph less accurate at small scales, it makes your graph *more* revealing at large scales."
+ "(\n",
+ " ggplot(penguins, aes(x=\"flipper_length_mm\", y=\"body_mass_g\"))\n",
+ " + geom_point(aes(color=\"species\", shape=\"island\"))\n",
+ ")"
]
},
{
"cell_type": "markdown",
- "id": "cd804cea",
+ "id": "14d0aacb",
"metadata": {},
"source": [
- "## Co-ordinates and Scales\n",
+ "However adding too many aesthetic mappings to a plot makes it cluttered and difficult to make sense of.\n",
"\n",
- "### Co-ordinates\n",
+ "Another way, which is particularly useful for categorical variables, is to split your plot into **facets** (also known as **small multiples**), subplots that each display one subset of the data.\n",
"\n",
- "The co-ordinates of a plot are the system that determines which data is attached to which axis of, typically, the horizontal, or x-axis, and the vertical, or y-axis. This is set by arguments to the call to `so.Plot`, so to reverse the plot from before we simply reverse the arguments:"
+ "To facet your plot by a single variable, use `facet_wrap()`.\n",
+ "\n",
+ "The first argument of `facet_wrap()` tells the function what variable to have in successive charts. The variable that you pass to `facet_wrap()` should be categorical."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "5c4be777",
+ "id": "00dd36e3",
"metadata": {},
"outputs": [],
"source": [
- "so.Plot(mpg, \"hwy\", \"displ\").add(so.Dot())"
+ "(\n",
+ " ggplot(penguins, aes(x=\"flipper_length_mm\", y=\"body_mass_g\"))\n",
+ " + geom_point(aes(color=\"species\", shape=\"species\"))\n",
+ " + facet_wrap(facets=\"island\")\n",
+ ")"
]
},
{
"cell_type": "markdown",
- "id": "cd10e1bf",
+ "id": "ee5a3eed",
"metadata": {},
"source": [
- "You can also do this explicitly by setting `x=\"hwy\"` and `y=\"displ\"`, and there's a lot to be said for being explicit (when you read your code back later, it's very helpful indeed)."
+ "You will learn about many other geoms for visualising distributions of variables and relationships between them in later chapters."
]
},
{
"cell_type": "markdown",
- "id": "77f45751",
+ "id": "c4566786",
"metadata": {},
"source": [
- "### Scales\n",
+ "### Exercises\n",
"\n",
- "Let's say you create a chart but the data vary on a scale that isn't shown well by the default axes. If you find yourself in this situation, you may wish to change the *scale* of one or both of the axes. This is controlled by the `Scale` property in **seaborn**.\n",
+ "1. Make a scatterplot of `bill_depth_mm` vs. `bill_length_mm` and color the points by `species`.\n",
+ " What does adding coloring by species reveal about the relationship between these two variables?\n",
+ " What about faceting by `species`?\n",
+ "\n",
+ "2. Why does the following yield two separate legends?\n",
+ " How would you fix it to combine the two legends?\n",
+ "\n",
+ " ```python\n",
+ " (\n",
+ " ggplot(\n",
+ " data = penguins,\n",
+ " mapping = aes(\n",
+ " x = \"bill_length_mm\", y = \"bill_depth_mm\", \n",
+ " color = \"species\", shape = \"species\"\n",
+ " )\n",
+ " ) +\n",
+ " geom_point() +\n",
+ " labs(color = \"Species\")\n",
+ " )\n",
+ " ```\n",
"\n",
- "The notion of scaling will probably not be unfamiliar; it means that a mathematical transformation, such as log, is made to the coordinate (or axes) variables.\n",
+ "3. Create the two following stacked bar plots.\n",
+ " Which question can you answer with the first one?\n",
+ " Which question can you answer with the second one?\n",
"\n",
- "We'll show this using the `planets` dataset, which has lots of variation in it!"
+ " ```python\n",
+ " ggplot(penguins, aes(x = \"island\", fill = \"species\")) +\n",
+ " geom_bar(position = \"fill\")\n",
+ " ggplot(penguins, aes(x = \"species\", fill = \"island\")) +\n",
+ " geom_bar(position = \"fill\")\n",
+ " ```\n"
]
},
{
- "cell_type": "code",
- "execution_count": null,
- "id": "20d36460",
+ "cell_type": "markdown",
+ "id": "150fbd1f",
"metadata": {},
- "outputs": [],
"source": [
- "planets = sns.load_dataset(\"planets\").query(\"distance < 1000\")\n",
- "planets.head()"
+ "## Saving your plots\n",
+ "\n",
+ "Once you've made a plot, you might want to save it as an image that you can use elsewhere.\n",
+ "That's the job of `ggsave()`, which will save the plot most recently created to disk:\n"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "fbed3933",
+ "id": "3410634b",
"metadata": {},
"outputs": [],
"source": [
- "(so.Plot(planets, x=\"mass\", y=\"distance\").scale(x=\"log\", y=\"log\").add(so.Dot()))"
+ "plotted_data = (\n",
+ " ggplot(penguins, aes(x=\"flipper_length_mm\", y=\"body_mass_g\")) + geom_point()\n",
+ ")\n",
+ "ggsave(plotted_data, filename=\"penguin-plot.svg\")"
]
},
{
"cell_type": "markdown",
- "id": "994e46ce",
+ "id": "ee6fa32c",
"metadata": {},
"source": [
- "Here we used a log scale for both the x- and y-axes because both mass and distance vary over many orders of magnitude.\n",
+ "This saved the figure to disk at the location shown—by default it's in a subdirectory called \"lets-plot-images\".\n",
+ "\n",
+ "We used the file format \"svg\". There are lots of output options to choose from to save your file to. Remember that, for graphics, *vector formats* are generally better than *raster formats*. In practice, this means saving plots in svg or pdf formats over jpg or png file formats. The svg format works in a lot of contexts (including Microsoft Word) and is a good default. To choose between formats, just supply the file extension and the file type will change automatically, eg \"chart.svg\" for svg or \"chart.png\" for png. You can also save figures in HTML format.\n",
"\n",
- "But the scale property can apply to other dimensions that we are visualising in our plots too; here's an example where we're using colour (in the below, plasma is the name of a built-in continuous colourmap, a way of representing a continuous number line with colour gradients):"
+ "If you're using a raster format then you'll need to specify how big the figure is via the *scale* keyword argument."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "8bbad703",
- "metadata": {},
+ "id": "852afe51",
+ "metadata": {
+ "tags": [
+ "remove-cell"
+ ]
+ },
"outputs": [],
"source": [
- "(\n",
- " so.Plot(planets, x=\"mass\", y=\"distance\", color=\"orbital_period\")\n",
- " .scale(x=\"log\", y=\"log\", color=so.Continuous(\"plasma\", trans=\"log\"))\n",
- " .add(so.Dot())\n",
- ")"
+ "import shutil\n",
+ "\n",
+ "shutil.rmtree(\"lets-plot-images\")"
]
},
{
"cell_type": "markdown",
- "id": "a7589df3",
+ "id": "2987bf18",
"metadata": {},
"source": [
- "Sometimes you *don't* want to apply the transform to everything, and that's okay too. Here's an example where the log scale *doesn't* apply to the mass variable (even though it's shown)."
+ "### Exercises\n",
+ "\n",
+ "1. Save the figure above as a PNG. Try varying the scale."
]
},
{
- "cell_type": "code",
- "execution_count": null,
- "id": "d3685f89",
+ "cell_type": "markdown",
+ "id": "1390edc2",
"metadata": {},
- "outputs": [],
"source": [
- "(\n",
- " so.Plot(planets, x=\"distance\", y=\"orbital_period\", pointsize=\"mass\")\n",
- " .scale(x=\"log\", y=\"log\", pointsize=None)\n",
- " .add(so.Dot())\n",
- ")"
+ "## Common Problems\n",
+ "\n",
+ "As you start to run code, you're likely to run into problems.\n",
+ "Don't worry—it happens to everyone.\n",
+ "We have all been writing Python code for years, but every day we still write code that doesn't work on the first try!\n",
+ "\n",
+ "Start by carefully comparing the code that you're running to the code in the book: A misplaced character can make all the difference!\n",
+ "Make sure that every `(` is matched with a `)` and every `\"` is paired with another `\"`. In Visual Studio Code, you can get extensions that colour match brackets so you can easily see if you closed them or not.\n",
+ "Sometimes you'll run the code and nothing happens.\n",
+ "\n",
+ "One common problem when creating **letsplot** graphics is to put the `+` in the wrong place: it has to come at the end of the line, not the start.\n",
+ "\n",
+ "\n",
+ "If you're still stuck, try the help.\n",
+ "You can get help about any Python function by running `help(function_name)` in the interactive window.\n",
+ "Don't worry if the help doesn't seem that helpful - instead skip down to the examples and look for code that matches what you're trying to do.\n",
+ "\n",
+ "If you're still stuck, check out the **letsplot** [documentation](https://lets-plot.org/) or doing a Google search (especially helpful for error messages).\n"
]
},
{
"cell_type": "markdown",
- "id": "a5a522b3",
+ "id": "f33dc022",
"metadata": {},
"source": [
- "## Summing Up\n",
+ "# Summary\n",
"\n",
- "In the above, you've got to grips with some of the basics of visualisation with **seaborn**. You can find much more information in the documentation for that project. But let's recap the grammar of a **seaborn** plot. The typical call will look something like this:\n",
+ "In this chapter, you've learned the basics of data visualisation with ggplot2.\n",
+ "We started with the basic idea that underpins **letsplot**: a visualisation is a mapping from variables in your data to aesthetic properties like position, colour, size and shape.\n",
+ "You then learned about increasing the complexity and improving the presentation of your plots layer-by-layer.\n",
+ "You also learned about commonly used plots for visualising the distribution of a single variable as well as for visualising relationships between two or more variables, by leveraging additional aesthetic mappings and/or splitting your plot into small multiples using faceting.\n",
"\n",
- "```python\n",
- "(\n",
- " so.Plot(, x=, y=, )\n",
- " .scale(x=, )\n",
- " .add(, )\n",
- " .add()\n",
- ")\n",
- "```"
+ "We'll use visualisations again and again throughout this book, introducing new techniques as we need them as well as do a deeper dive into creating visualisations with **letsplot** in subsequent chapters.\n",
+ "\n",
+ "With the basics of visualisation under your belt, in the next chapter we're going to switch gears a little and give you some practical workflow advice.\n",
+ "We intersperse workflow advice with data science tools throughout this part of the book because it'll help you stay organised as you write more Python code."
]
}
],
@@ -891,7 +1126,7 @@
"name": "python",
"nbconvert_exporter": "python",
"pygments_lexer": "ipython3",
- "version": "3.9.12"
+ "version": "3.10.12"
},
"toc-showtags": true
},
diff --git a/databases.ipynb b/databases.ipynb
index 30747e0..fe7dda5 100644
--- a/databases.ipynb
+++ b/databases.ipynb
@@ -493,7 +493,7 @@
"source": [
"track.group_by(\"AlbumId\").mutate(\n",
" mean_mins_track=track.Milliseconds.mean() / 1e3 / 60\n",
- ").sort_by(\"mean_mins_track\").limit(5)"
+ ").order_by(\"mean_mins_track\").limit(5)"
]
},
{
@@ -802,7 +802,7 @@
"name": "python",
"nbconvert_exporter": "python",
"pygments_lexer": "ipython3",
- "version": "3.9.12"
+ "version": "3.10.12"
},
"toc-showtags": true,
"vscode": {
diff --git a/environment.yml b/environment.yml
index 77c83ff..52d5278 100644
--- a/environment.yml
+++ b/environment.yml
@@ -1,5 +1,5 @@
-name: python4DS
+name: py4ds2e
channels:
- conda-forge
dependencies:
@@ -7,7 +7,7 @@ dependencies:
- numpy
- pandas
- pip
- - python=3.9
+ - python=3.10
- pyyaml
- scipy
- statsmodels
@@ -19,25 +19,24 @@ dependencies:
- jupyter-book
- pytest
- pre-commit
- - linearmodels
- - pingouin
- - plotly
- - plotnine
- - altair
- jupyterlab
- nbstripout
- - vega_datasets
- ghp-import
- pip
- black
- black-jupyter
- beautifulsoup4
+ - geopandas
- pip:
- skimpy
- pyarrow
- - pandas-profiling
- watermark
- graphviz
- openpyxl
- sqlmodel
- - ibis-framework
\ No newline at end of file
+ - ibis-framework
+ - lets-plot
+ - polars
+ - palmerpenguins
+ - pandas-profiling
+ - rich
\ No newline at end of file
diff --git a/exploratory-data-analysis.ipynb b/exploratory-data-analysis.ipynb
index 4f71ce8..26c1545 100644
--- a/exploratory-data-analysis.ipynb
+++ b/exploratory-data-analysis.ipynb
@@ -22,7 +22,26 @@
"\n",
"### Prerequisites\n",
"\n",
- "For doing EDA, we'll use the **pandas**, **skimpy**, and **pandas-profiling** packages. You are likely to already have **pandas** installed. We'll also need **seaborn** for data visualisation, which can you install with `pip install --pre seaborn`. To install the other two packages, open up a terminal in Visual Studio Code and run `pip install skimpy` and `pip install pandas-profiling`."
+ "For doing EDA, we'll use the **pandas**, **skimpy**, and **pandas-profiling** packages. We'll also need **lets-plot** for data visualisation. All of these can be installed via `pip install `.\n",
+ "\n",
+ "As ever, we begin by loading these packages that we'll use:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "a3377aa6",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from skimpy import skim\n",
+ "from pandas_profiling import ProfileReport\n",
+ "import pandas as pd\n",
+ "from pandas.api.types import CategoricalDtype\n",
+ "from lets_plot import *\n",
+ "from lets_plot.mapping import as_discrete\n",
+ "\n",
+ "LetsPlot.setup_html()"
]
},
{
@@ -52,29 +71,23 @@
"## Questions\n",
"\n",
"> \"There are no routine statistical questions, only questionable statistical routines.\" --- Sir David Cox\n",
+ "\n",
"> \"Far better an approximate answer to the right question, which is often vague, than an exact answer to the wrong question, which can always be made precise.\" --- John Tukey\n",
"\n",
"Your goal during EDA is to develop an understanding of your data. The easiest way to do this is to use questions as tools to guide your investigation. When you ask a question, the question focuses your attention on a specific part of your dataset and helps you decide which graphs, models, or transformations to make.\n",
"\n",
"EDA is fundamentally a creative process. And like most creative processes, the key to asking *quality* questions is to generate a large *quantity* of questions. It is difficult to ask revealing questions at the start of your analysis because you do not know what insights are contained in your dataset. On the other hand, each new question that you ask will expose you to a new aspect of your data and increase your chance of making a discovery. You can quickly drill down into the most interesting parts of your data---and develop a set of thought-provoking questions---if you follow up each question with a new question based on what you find.\n",
"\n",
- "There is no rule about which questions you should ask to guide your research. However, two types of questions will always be useful for making discoveries within your data. You can loosely word these questions as:\n",
+ "There is no rule about which questions you should ask to guide your research.\n",
+ "However, two types of questions will always be useful for making discoveries within your data.\n",
+ "You can loosely word these questions as:\n",
"\n",
"1. What type of variation occurs within my variables?\n",
"\n",
"2. What type of covariation occurs between my variables?\n",
"\n",
- "The rest of this chapter will look at these two questions. We'll explain what variation and covariation are, and We'll show you several ways to answer each question. To make the discussion easier, let's define some terms:\n",
- "\n",
- "- A **variable** is a quantity, quality, or property that you can measure.\n",
- "\n",
- "- A **value** is the state of a variable when you measure it. The value of a variable may change from measurement to measurement.\n",
- "\n",
- "- An **observation** is a set of measurements made under similar conditions (you usually make all of the measurements in an observation at the same time and on the same object). An observation will contain several values, each associated with a different variable. We'll sometimes refer to an observation as a data point.\n",
- "\n",
- "- **Tabular data** is a set of values, each associated with a variable and an observation. Tabular data is *tidy* if each value is placed in its own \"cell\", each variable in its own column, and each observation in its own row.\n",
- "\n",
- "So far, all of the data that you've seen has been tidy. In real-life, most data isn't tidy, so we'll come back to how to clean untidy data later in the book."
+ "The rest of this chapter will look at these two questions.\n",
+ "We'll explain what variation and covariation are, and we'll show you several ways to answer each question."
]
},
{
@@ -84,290 +97,613 @@
"source": [
"## Variation\n",
"\n",
- "**Variation** is the tendency of the values of a variable to change from measurement to measurement. You can see variation easily in real life; if you measure any continuous variable twice, you will get two different results. This is true even if you measure quantities that are constant, like the speed of light. Each of your measurements include a small amount of error that varies from measurement to measurement. Variables can also vary if you measure across different subjects (e.g. the eye colours of different people) or different times (e.g. the energy levels of an electron at different moments).\n",
- "\n",
- "Every variable has its own pattern of variation, which can reveal interesting information about how that variable varies between measurements on the same observation as well as across observations. The best way to understand that pattern is to visualise the distribution of the variable's values.\n",
- "\n",
- "### Visualising distributions\n",
+ "**Variation** is the tendency of the values of a variable to change from measurement to measurement. You can see variation easily in real life; if you measure any continuous variable twice, you will get two different results. This is true even if you measure quantities that are constant, like the speed of light, because of deficiencies in equipment. Each of your measurements includes a small amount of error that varies from measurement to measurement. Variables can also vary if you measure across different subjects (e.g. the eye colours of different people) or different times (e.g. the energy levels of an electron at different moments). The best way to understand that pattern is to visualize the distribution of the variable's values.\n",
"\n",
- "How you visualize the distribution of a variable will depend on whether the variable is categorical or continuous. A variable is **categorical** if it can only take one of a small set of values. In data analysis in Python, categorical variables are usually saved as the 'category' type in **pandas** data frames. To examine the distribution of a categorical variable, you can use a bar chart. First let's load up **seaborn** for visusalisation and load up the diamonds dataset in **pandas**."
+ "We'll start our exploration by visualizing the distribution of weights (`\"carat\"`) of \\~54,000 diamonds from the `diamonds` dataset."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "75e3b802",
+ "id": "069caa7c",
"metadata": {},
"outputs": [],
"source": [
- "import seaborn.objects as so"
+ "diamonds = pd.read_csv(\n",
+ " \"https://github.com/mwaskom/seaborn-data/raw/master/diamonds.csv\"\n",
+ ")\n",
+ "diamonds[\"cut\"] = diamonds[\"cut\"].astype(\n",
+ " CategoricalDtype(\n",
+ " categories=[\"Fair\", \"Good\", \"Very Good\", \"Premium\", \"Ideal\"], ordered=True\n",
+ " )\n",
+ ")\n",
+ "diamonds[\"color\"] = diamonds[\"color\"].astype(\n",
+ " CategoricalDtype(categories=[\"D\", \"E\", \"F\", \"G\", \"H\", \"I\", \"J\"], ordered=True)\n",
+ ")\n",
+ "diamonds.head()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "01f5979c",
+ "metadata": {},
+ "source": [
+ "Since `\"carat\"` is a numerical variable, we can use a histogram:"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "e7f4de95",
+ "id": "97900f58",
"metadata": {},
"outputs": [],
"source": [
- "import pandas as pd\n",
- "\n",
- "diamonds = pd.read_csv(\n",
- " \"https://github.com/mwaskom/seaborn-data/raw/master/diamonds.csv\"\n",
- ")\n",
- "diamonds[\"cut\"] = diamonds[\"cut\"].astype(\"category\")\n",
- "diamonds.head()"
+ "(ggplot(diamonds, aes(x=\"carat\")) + geom_histogram(binwidth=0.5))"
]
},
{
"cell_type": "markdown",
- "id": "86240820",
+ "id": "2307ba7c",
"metadata": {},
"source": [
- "Now we can visualise the data using a bar chart:"
+ "Now that you can visualize variation, what should you look for in your plots?\n",
+ "And what type of follow-up questions should you ask?\n",
+ "We've put together a list below of the most useful types of information that you will find in your graphs, along with some follow-up questions for each type of information.\n",
+ "The key to asking good follow-up questions will be to rely on your curiosity (What do you want to learn more about?) as well as your skepticism (How could this be misleading?).\n",
+ "\n",
+ "### Typical values\n",
+ "\n",
+ "In both bar charts and histograms, tall bars show the common values of a variable, and shorter bars show less-common values.\n",
+ "Places that do not have bars reveal values that were not seen in your data.\n",
+ "To turn this information into useful questions, look for anything unexpected:\n",
+ "\n",
+ "- Which values are the most common?\n",
+ " Why?\n",
+ "\n",
+ "- Which values are rare?\n",
+ " Why?\n",
+ " Does that match your expectations?\n",
+ "\n",
+ "- Can you see any unusual patterns?\n",
+ " What might explain them?\n",
+ "\n",
+ "Let's take a look at the distribution of `\"carat\"` for smaller diamonds.\n",
+ "\n",
+ "Note that when we create `smaller_diamonds`, we're doing it by creating a copy. Otherwise, any changes we made to `smaller_diamonds` would also affect `diamonds` (the two point to the same underlying data in your computer's memory). Sometimes you may want to have a cut still be connected to the original dataset, and sometimes you don't; in this case, we'd like them to be distinct so we use `copy()`."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "43eca1b9",
+ "id": "20d75550",
"metadata": {},
"outputs": [],
"source": [
- "(so.Plot(diamonds, \"cut\").add(so.Bar(), so.Hist()))"
+ "smaller_diamonds = diamonds.query(\"carat < 3\").copy()\n",
+ "\n",
+ "(ggplot(smaller_diamonds, aes(x=\"carat\")) + geom_histogram(binwidth=0.01))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "ba20a0e2",
+ "metadata": {},
+ "source": [
+ "This histogram suggests several interesting questions:\n",
+ "\n",
+ "- Why are there more diamonds at whole carats and common fractions of carats?\n",
+ "\n",
+ "- Why are there more diamonds slightly to the right of each peak than there are slightly to the left of each peak?\n",
+ "\n",
+ "Visualisations can also reveal clusters, which suggest that subgroups exist in your data.\n",
+ "To understand the subgroups, ask:\n",
+ "\n",
+ "- How are the observations within each subgroup similar to each other?\n",
+ "\n",
+ "- How are the observations in separate clusters different from each other?\n",
+ "\n",
+ "- How can you explain or describe the clusters?\n",
+ "\n",
+ "- Why might the appearance of clusters be misleading?\n",
+ "\n",
+ "Some of these questions can be answered with the data while some will require domain expertise about the data.\n",
+ "Many of them will prompt you to explore a relationship *between* variables, for example, to see if the values of one variable can explain the behavior of another variable.\n",
+ "We'll get to that shortly."
]
},
{
"cell_type": "markdown",
- "id": "eb60dea5",
+ "id": "0626d35a",
"metadata": {},
"source": [
- "The height of the bars displays how many observations occurred with each x value. You can compute these values directly with **pandas** too:"
+ "### Unusual values\n",
+ "\n",
+ "Outliers are observations that are unusual; data points that don't seem to fit the pattern.\n",
+ "Sometimes outliers are data entry errors, sometimes they are simply values at the extremes that happened to be observed in this data collection, and other times they suggest important new discoveries.\n",
+ "When you have a lot of data, outliers are sometimes difficult to see in a histogram.\n",
+ "For example, take the distribution of the `\"y\"` variable from the diamonds dataset.\n",
+ "The only evidence of outliers is the unusually wide limits on the x-axis.\n"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "934db5b6",
+ "id": "d9d7e995",
"metadata": {},
"outputs": [],
"source": [
- "diamonds[\"cut\"].value_counts()"
+ "(ggplot(diamonds, aes(x=\"y\")) + geom_histogram(binwidth=0.5))"
]
},
{
"cell_type": "markdown",
- "id": "953fd423",
+ "id": "05bcf733",
"metadata": {},
"source": [
- "A variable is **continuous** if it can take any of an infinite set of ordered values. Numbers and date-times are two examples of continuous variables. To examine the distribution of a continuous variable, you can use a histogram:"
+ "There are so many observations in the common bins that the rare bins are very short, making it very difficult to see them (although maybe if you stare intently at 0 you'll spot something).\n",
+ "To make it easy to see the unusual values, we need to zoom to small values of the y-axis with `coord_cartesian()`:"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "f5c4cc5d",
+ "id": "ea8f8bf3",
"metadata": {},
"outputs": [],
"source": [
- "(so.Plot(diamonds, \"carat\").add(so.Bar(), so.Hist(binwidth=0.5)))"
+ "(\n",
+ " ggplot(diamonds, aes(x=\"y\"))\n",
+ " + geom_histogram(binwidth=0.5)\n",
+ " + coord_cartesian(ylim=[0, 50])\n",
+ ")"
]
},
{
"cell_type": "markdown",
- "id": "fc08384d",
+ "id": "ba2b2c79",
"metadata": {},
"source": [
- "You can also compute this directly using **pandas** using `pd.cut` to assign a category (an interval) to each row and then `value_counts()` to count the number of rows in each category."
+ "`coord_cartesian()` also has an `xlim()` argument for when you need to zoom into the x-axis.\n",
+ "**Lets-Plot** also has `xlim()` and `ylim()` functions that work slightly differently: they throw away the data outside the limits.\n",
+ "\n",
+ "This allows us to see that there are three unusual values: 0, \\~30, and \\~60. We pluck them out with **pandas**:"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "03d87bcb",
+ "id": "e81ffb55",
"metadata": {},
"outputs": [],
"source": [
- "pd.cut(diamonds[\"carat\"], bins=11).value_counts()"
+ "unusual = diamonds.query(\"y < 3 or y > 20\").loc[:, [\"x\", \"y\", \"z\", \"price\"]]\n",
+ "unusual"
]
},
{
"cell_type": "markdown",
- "id": "d9e9b67d",
+ "id": "c9321f36",
"metadata": {},
"source": [
- "A histogram divides the x-axis into equally spaced bins and then uses the height of a bar to display the number of observations that fall in each bin. You can set the number of intervals in a histogram plot with the `binwidth=` keyword argument, which is measured in the units of the `x` variable. You should always explore a variety of binwidths when working with histograms, as different binwidths can reveal different patterns.\n",
+ "The `\"y\"` variable measures one of the three dimensions of these diamonds, in mm.\n",
+ "We know that diamonds can't have a width of 0mm, so these values must be incorrect.\n",
+ "By doing EDA, we have discovered missing data that was coded as 0, which we never would have found by simply searching for `NA`s.\n",
+ "Going forward we might choose to re-code these values as `NA`s in order to prevent misleading calculations.\n",
+ "We might also suspect that measurements of 32mm and 59mm are implausible: those diamonds are over an inch long, but don't cost hundreds of thousands of dollars!\n",
"\n",
- "For example, here is how the graph above looks when we zoom into just the diamonds with a size of less than three carats and choose a smaller binwidth.\n"
+ "It's good practice to repeat your analysis with and without the outliers.\n",
+ "If they have minimal effect on the results, and you can't figure out why they're there, it's reasonable to omit them, and move on.\n",
+ "However, if they have a substantial effect on your results, you shouldn't drop them without justification.\n",
+ "You'll need to figure out what caused them (e.g., a data entry error) and disclose that you removed them in your write-up.\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "142d21d7",
+ "metadata": {},
+ "source": [
+ "### Exercises\n",
+ "\n",
+ "1. Explore the distribution of each of the `x`, `y`, and `z` variables in `diamonds`.\n",
+ " What do you learn?\n",
+ " Think about a diamond and how you might decide which dimension is the length, width, and depth.\n",
+ "\n",
+ "2. Explore the distribution of `\"price\"`.\n",
+ " Do you discover anything unusual or surprising?\n",
+ " (Hint: Carefully think about the `binwidth=` keyword argument setting and make sure you try a wide range of values.)\n",
+ "\n",
+ "3. How many diamonds are 0.99 carat?\n",
+ " How many are 1 carat?\n",
+ " What do you think is the cause of the difference?\n",
+ "\n",
+ "4. Compare and contrast `coord_cartesian()` vs. `xlim()` or `ylim()` when zooming in on a histogram.\n",
+ " What happens if you leave binwidth unset?\n",
+ " What happens if you try and zoom so only half a bar shows?\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "c51e8d6f",
+ "metadata": {},
+ "source": [
+ "## Unusual Values\n",
+ "\n",
+ "If you've encountered unusual values in your dataset, and simply want to move on to the rest of your analysis, you have two options.\n",
+ "\n",
+ "1. Drop the entire row with the strange values:\n",
+ "\n",
+ " ```python\n",
+ " condition = ((diamonds[\"y\"] < 3) | (diamonds[\"y\"] > 20))\n",
+ " diamonds2 = diamonds.loc[~condition, :]\n",
+ " ```\n",
+ "\n",
+ " We don't recommend this option because one invalid value doesn't imply that all the other values for that observation are also invalid.\n",
+ " Additionally, if you have low quality data, by the time that you've applied this approach to every variable you might find that you don't have any data left!\n",
+ "\n",
+ "2. Instead, we recommend replacing the unusual values with missing values.\n",
+ " One way to do this, which makes a distinction between dataframes that have had the unusual values replaced and the original data, is to make a copy and then set the problematic values to `pd.NA`, **pandas**'s special NA value.\n"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "7a142d69",
+ "id": "ecf345a7",
"metadata": {},
"outputs": [],
"source": [
- "(so.Plot(diamonds.query(\"carat < 3\"), \"carat\").add(so.Bar(), so.Hist(binwidth=0.1)))"
+ "diamonds2 = diamonds.copy()\n",
+ "condition = (diamonds2[\"y\"] < 3) | (diamonds2[\"y\"] > 20)\n",
+ "diamonds2.loc[condition, \"y\"] = pd.NA"
]
},
{
"cell_type": "markdown",
- "id": "9a8f6dbe",
+ "id": "d26d922e",
"metadata": {},
"source": [
- "Now that you can visualise variation, what should you look for in your plots? And what type of follow-up questions should you ask? Below is a list of the most useful types of information that you will find in your graphs, along with some follow-up questions for each type of information. The key to asking good follow-up questions will be to rely on your curiosity (What do you want to learn more about?) as well as your skepticism (How could this be misleading?)."
+ "It's not obvious where you should plot missing values, so **lets-plot** doesn't include them in the plot:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "15a43255",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(diamonds2, aes(x=\"x\", y=\"y\")) + geom_point())"
]
},
{
"cell_type": "markdown",
- "id": "0cc70e63",
+ "id": "e8ae854b",
+ "metadata": {},
+ "source": [
+ "Other times you want to understand what makes observations with missing values different to observations with recorded values.\n",
+ "For example, in the nycflights13 data, missing values in the `\"dep_time\"` variable indicate that the flight was cancelled.\n",
+ "So you might want to compare the scheduled departure times for cancelled and non-cancelled times.\n",
+ "You can do this by making a new variable, using `is.na()` to check if `\"dep_time\"` is missing."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "0a4ea922",
"metadata": {},
+ "outputs": [],
"source": [
- "### Typical Values\n",
+ "url = \"https://raw.githubusercontent.com/byuidatascience/data4python4ds/master/data-raw/flights/flights.csv\"\n",
+ "flights = pd.read_csv(url)\n",
+ "flights.head()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "6849f4d9",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "flights2 = flights.assign(\n",
+ " cancelled=lambda x: pd.isna(x[\"dep_time\"]),\n",
+ " sched_hour=lambda x: x[\"sched_dep_time\"] // 100,\n",
+ " sched_min=lambda x: x[\"sched_dep_time\"] % 100,\n",
+ " sched_dep_time=lambda x: x[\"sched_hour\"] + x[\"sched_min\"] / 60,\n",
+ ")\n",
"\n",
- "In both bar charts and histograms, tall bars show the common values of a variable, and shorter bars show less-common values.\n",
- "Places that do not have bars reveal values that were not seen in your data.\n",
- "To turn this information into useful questions, look for anything unexpected:\n",
+ "(\n",
+ " ggplot(flights2, aes(x=\"sched_dep_time\"))\n",
+ " + geom_freqpoly(aes(color=\"cancelled\"), binwidth=1 / 4)\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "b97e453a",
+ "metadata": {},
+ "source": [
+ "However this plot isn't great because there are many more non-cancelled flights than cancelled flights.\n",
+ "In the next section we'll explore some techniques for improving this comparison.\n",
"\n",
- "- Which values are the most common?\n",
- " Why?\n",
+ "### Exercises\n",
"\n",
- "- Which values are rare?\n",
- " Why?\n",
- " Does that match your expectations?\n",
+ "1. What happens to missing values in a histogram?\n",
+ " What happens to missing values in a bar chart?\n",
+ " Why is there a difference in how missing values are handled in histograms and bar charts?\n",
"\n",
- "- Can you see any unusual patterns?\n",
- " What might explain them?\n",
+ "2. Recreate the frequency plot of `scheduled_dep_time` coloured by whether the flight was cancelled or not.\n",
+ " Also facet by the `cancelled` variable.\n",
+ " Experiment with different values of the `scales` variable in the faceting function to mitigate the effect of more non-cancelled flights than cancelled flights.\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "f63d0b9f",
+ "metadata": {},
+ "source": [
+ "## Covariation\n",
"\n",
- "As an example, the histogram below suggests several interesting questions:\n",
+ "If variation describes the behavior *within* a variable, covariation describes the behavior *between* variables.\n",
+ "**Covariation** is the tendency for the values of two or more variables to vary together in a related way.\n",
+ "The best way to spot covariation is to visualise the relationship between two or more variables, but note that covariation doesn't imply a causal relationship between variables.\n",
"\n",
- "- Why are there more diamonds at whole carats and common fractions of carats?\n",
+ "### A categorical and a numerical variable\n",
"\n",
- "- Why are there more diamonds slightly to the right of each peak than there are slightly to the left of each peak?"
+ "For example, let's explore how the price of a diamond varies with its quality (measured by `\"cut\"`) using `geom_freqpoly()`:\n"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "b131f1ba",
+ "id": "e1719d8f",
"metadata": {},
"outputs": [],
"source": [
- "(so.Plot(diamonds.query(\"carat < 3\"), \"carat\").add(so.Bar(), so.Hist(binwidth=0.01)))"
+ "(\n",
+ " ggplot(diamonds, aes(x=\"price\"))\n",
+ " + geom_freqpoly(aes(color=\"cut\"), binwidth=500, linewidth=0.75)\n",
+ ")"
]
},
{
"cell_type": "markdown",
- "id": "4a6b18aa",
+ "id": "20387b96",
"metadata": {},
"source": [
- "Clusters of similar values suggest that subgroups exist in your data.\n",
- "To understand the subgroups, ask:\n",
+ "The default appearance of `geom_freqpoly()` is not that useful here because the height, determined by the overall count, differs so much across cuts, making it hard to see the differences in the shapes of their distributions.\n",
"\n",
- "- How are the observations within each cluster similar to each other?\n",
+ "To make the comparison easier we need to swap what is displayed on the y-axis.\n",
+ "Instead of displaying count, we'll display the **density**, which is the count standardised so that the area under each frequency polygon is one."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "9388e24b",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(\n",
+ " ggplot(diamonds, aes(x=\"price\"))\n",
+ " + geom_density(aes(color=\"cut\", fill=\"cut\"), size=1, alpha=0.2)\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "157f63dd",
+ "metadata": {},
+ "source": [
+ "There’s something rather surprising about this plot - it appears that fair diamonds (the lowest quality) have the highest average price! But maybe that’s because density plots are a little hard to interpret - there’s a lot going on in this plot.\n",
"\n",
- "- How are the observations in separate clusters different from each other?\n",
+ "A visually simpler plot for exploring this relationship is using side-by-side boxplots."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "a3f333a6",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(diamonds, aes(x=\"cut\", y=\"price\")) + geom_boxplot())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "b9d9ef00",
+ "metadata": {},
+ "source": [
+ "We see much less information about the distribution, but the boxplots are much more compact so we can more easily compare them (and fit more on one plot). It supports the counter-intuitive finding that better quality diamonds are typically cheaper! In the exercises, you’ll be challenged to figure out why.\n",
"\n",
- "- How can you explain or describe the clusters?\n",
+ "`\"cut\"` is an ordered categorical variable: fair is worse than good, which is worse than very good and so on. Many categorical variables don’t have such an intrinsic order, so you might want to reorder them to make a more informative display. One way to do that is according to the median value, though other options are available.\n",
"\n",
- "- Why might the appearance of clusters be misleading?\n",
+ "As an example, with the mpg dataset, we might want to look at how highway mileage varies across classes:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "6949db81",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "mpg = pd.read_csv(\n",
+ " \"https://vincentarelbundock.github.io/Rdatasets/csv/ggplot2/mpg.csv\", index_col=0\n",
+ ")\n",
+ "mpg[\"class\"] = mpg[\"class\"].astype(\"category\")\n",
"\n",
- "Many of the questions above will prompt you to explore a relationship *between* variables, for example, to see if the values of one variable can explain the behavior of another variable. We'll get to that shortly."
+ "(ggplot(mpg, aes(x=\"class\", y=\"hwy\")) + geom_boxplot())"
]
},
{
"cell_type": "markdown",
- "id": "29461679",
+ "id": "871aaf1c",
"metadata": {},
"source": [
- "### Unusual Values\n",
- "\n",
- "Outliers are observations that are unusual; data points that don't seem to fit the pattern.\n",
- "Sometimes outliers are data entry errors; other times outliers suggest important new science.\n",
- "When you have a lot of data, outliers are sometimes difficult to see in a histogram.\n",
- "For example, take the distribution of the `y` variable from the diamonds dataset.\n",
- "The only evidence of outliers is the unusually wide limits on the x-axis."
+ "To make the trend easier to see, we can reorder class based on the median value of `\"hwy\"`:"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "274112a0",
+ "id": "a5b1ed09",
"metadata": {},
"outputs": [],
"source": [
- "(so.Plot(diamonds, \"y\").add(so.Bar(), so.Hist(binwidth=0.5)))"
+ "(ggplot(mpg) + geom_boxplot(aes(as_discrete(\"class\", order_by=\"..middle..\"), \"hwy\")))"
]
},
{
"cell_type": "markdown",
- "id": "4702d7fc",
+ "id": "dde59236",
"metadata": {},
"source": [
- "There are so many observations in the common bins that the rare bins are very short, making it very difficult to see them (although maybe if you stare intently at 0 you'll spot something). To make it easy to see the unusual values, we need to zoom to small values of the y-axis with a special value of the axis limits:"
+ "If you have long variable names, geom_boxplot() will work better if you flip it 90°. You can do that by adding `coord_flip()`."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "06924e4c",
+ "id": "920a4268",
"metadata": {},
"outputs": [],
"source": [
"(\n",
- " so.Plot(diamonds, x=\"y\")\n",
- " .add(so.Bar(), so.Hist(binwidth=0.5))\n",
- " .limit(y=(0, 10), x=(-5, 60))\n",
+ " ggplot(mpg)\n",
+ " + geom_boxplot(aes(as_discrete(\"class\", order_by=\"..middle..\"), \"hwy\"))\n",
+ " + coord_flip()\n",
")"
]
},
{
"cell_type": "markdown",
- "id": "b4bba593",
+ "id": "299635df",
"metadata": {},
"source": [
- "The `y` variable measures one of the three dimensions of these diamonds, in mm.\n",
- "We know that diamonds can't have a width of 0mm, so these values must be incorrect.\n",
- "We might also suspect that measurements of 32mm and 59mm are implausible: those diamonds are over an inch long, but don't cost hundreds of thousands of dollars!\n",
+ "#### Exercises\n",
"\n",
- "It's good practice to repeat your analysis with and without the outliers. If they have minimal effect on the results, and you can't figure out why they're there, it's reasonable to omit them, and move on.\n",
+ "1. Use what you've learned to improve the visualisation of the departure times of cancelled vs. non-cancelled flights.\n",
"\n",
- "However, if they have a substantial effect on your results, you shouldn't drop them without justification.\n",
- "You'll need to figure out what caused them (e.g. a data entry error) and disclose that you removed them in your write-up."
+ "2. Based on EDA, what variable in the diamonds dataset appears to be most important for predicting the price of a diamond?\n",
+ " How is that variable correlated with cut?\n",
+ " Why does the combination of those two relationships lead to lower quality diamonds being more expensive?\n",
+ "\n",
+ "3. Create a visualisation of diamond prices vs. a categorical variable from the `diamonds` dataset using `geom_violin()`, then a faceted `geom_histogram()`, then a coloured `geom_freqpoly()`, and then a coloured `geom_density()`.\n",
+ " Compare and contrast the four plots.\n",
+ " What are the pros and cons of each method of visualising the distribution of a numerical variable based on the levels of a categorical variable?\n",
+ "\n",
+ "4. If you have a small dataset, it's sometimes useful to use `geom_jitter()` to avoid overplotting to more easily see the relationship between a continuous and categorical variable.\n",
+ " The ggbeeswarm package provides a number of methods similar to `geom_jitter()`.\n",
+ " List them and briefly describe what each one does."
]
},
{
"cell_type": "markdown",
- "id": "4a0e2e67",
+ "id": "6e6a0b46",
"metadata": {},
"source": [
- "### Replacing Unusual Values\n",
+ "### Two categorical variables\n",
"\n",
- "If you’ve encountered unusual values in your dataset, and simply want to move on to the rest of your analysis (not your only response—you can also consider filling in the missing data), you have two options:\n",
+ "To visualise the covariation between categorical variables, you'll need to count the number of observations for each combination of levels of these categorical variables. You can do this with a `pd.crosstab` that we then melt to put it in \"tidy\" format."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "68d330d2",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ct_cut_color = pd.melt(\n",
+ " pd.crosstab(diamonds[\"cut\"], diamonds[\"color\"]).reset_index(),\n",
+ " id_vars=[\"cut\"],\n",
+ " value_vars=diamonds[\"color\"].unique(),\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "c2ebc0c6",
+ "metadata": {},
+ "source": [
+ "Followed by visualising it with `geom_tile`:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "e858cd22",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(ct_cut_color, aes(x=\"color\", y=\"cut\")) + geom_tile(aes(fill=\"value\")))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "8ccd5c1b",
+ "metadata": {},
+ "source": [
+ "### Exercises\n",
+ "\n",
+ "1. How could you rescale the count dataset above to more clearly show the distribution of cut within color, or colour within cut?\n",
"\n",
- "1. Drop the entire row with the strange values. You can do this by just working with a subset of the data, eg`diamonds.query('3 <= y <= 20')`. This option isn't generally recommended though as just because one measurement is invalid, it doesn’t mean all the measurements are. Additionally, if you have low quality data, by time that you’ve applied this approach to every variable you might find that you don’t have any data left!\n",
- "2. Replacing the unusual values with empty cells (ie remove those values entirely). The easiest way to do this is to use `assign()` to replace the variable with a modified copy. You can use the `np.where()` function to replace unusual values with `np.nan`, the **numpy** missing value operator:\n"
+ "2. What different data insights do you get with a segmented bar chart if colour is mapped to the x aesthetic and cut is mapped to the fill aesthetic? Calculate the counts that fall into each of the segments.\n",
+ "\n",
+ "3. Use `geom_tile()` together with **pandas** to explore how average flight departure delays vary by destination and month of year. What makes the plot difficult to read? How could you improve it?"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "b2f19afc",
+ "metadata": {},
+ "source": [
+ "### Two numerical variables\n",
+ "\n",
+ "You've already seen one great way to visualise the covariation between two numerical variables: draw a scatterplot with `geom_point()`.\n",
+ "You can see covariation as a pattern in the points.\n",
+ "For example, you can see a positive association between the carat size and price of a diamond: diamonds with more carats have a higher price.\n",
+ "The relationship is exponential."
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "d186896d",
+ "id": "2afe2535",
"metadata": {},
"outputs": [],
"source": [
- "import numpy as np\n",
+ "(ggplot(smaller_diamonds, aes(x=\"carat\", y=\"price\")) + geom_point())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "9d8034f2",
+ "metadata": {},
+ "source": [
+ "(In this section we'll use the `smaller_diamonds` dataset to stay focused on the bulk of the diamonds that are smaller than 3 carats)\n",
"\n",
- "diamonds[\"y\"] = diamonds[\"y\"].apply(lambda y: np.where(20 > y > 3, y, np.nan))"
+ "Scatterplots become less useful as the size of your dataset grows, because points begin to overplot, and pile up into areas of uniform black, making it hard to judge differences in the density of the data across the 2-dimensional space as well as making it hard to spot the trend.\n",
+ "You've already seen one way to fix the problem: using the `alpha` aesthetic to add transparency.\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "b55707a9",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(smaller_diamonds, aes(x=\"carat\", y=\"price\")) + geom_point(alpha=1 / 20))"
]
},
{
"cell_type": "markdown",
- "id": "53e73854",
+ "id": "351c22bd",
"metadata": {},
"source": [
- "`np.where()` typically has three arguments. The first argument condition should be a column of booleans. If `True`, then the next argument will be used; if `False`, the third. So we get the pattern `np.where(, , )`."
+ "But using transparency can be challenging for very large datasets. In that case, we recommend a *binscatter*, or binned scatterplot. A binned scatterplot divides the conditioning variable, `\"carat\"` in our example, into equally sized bins or quantiles, and then plots the conditional mean of the dependent variable, `\"price\"` in our example, within each bin. Bin scatters often come with confidence intervals too. A good bin scatter package in Python is [**binsreg**](https://nppackages.github.io/binsreg/). However, bin scatters are an advanced topic, and we won't cover them here."
]
},
{
@@ -452,8 +788,8 @@
"outputs": [],
"source": [
"(\n",
- " diamonds.groupby([\"cut\", \"color\"])\n",
- " .mean()[\"price\"]\n",
+ " diamonds.groupby([\"cut\", \"color\"])[\"price\"]\n",
+ " .mean()\n",
" .unstack()\n",
" .apply(lambda x: x / 1e3)\n",
" .fillna(\"-\")\n",
@@ -559,12 +895,12 @@
"source": [
"taxis = pd.read_csv(\"https://github.com/mwaskom/seaborn-data/raw/master/taxis.csv\")\n",
"# turn the pickup time column into a datetime\n",
- "# taxis[\"pickup\"] = pd.to_datetime(taxis[\"pickup\"])\n",
+ "taxis[\"pickup\"] = pd.to_datetime(taxis[\"pickup\"])\n",
"# set some other columns types\n",
"taxis = taxis.astype(\n",
" {\n",
- " \"dropoff\": \"datetime64\",\n",
- " \"pickup\": \"datetime64\",\n",
+ " \"dropoff\": \"datetime64[ns]\",\n",
+ " \"pickup\": \"datetime64[ns]\",\n",
" \"color\": \"category\",\n",
" \"payment\": \"category\",\n",
" \"pickup_zone\": \"string\",\n",
@@ -742,8 +1078,6 @@
"metadata": {},
"outputs": [],
"source": [
- "from skimpy import skim\n",
- "\n",
"skim(taxis)"
]
},
@@ -766,9 +1100,6 @@
"metadata": {},
"outputs": [],
"source": [
- "from pandas_profiling import ProfileReport\n",
- "\n",
- "\n",
"profile = ProfileReport(taxis, minimal=True, title=\"Profiling Report: Taxis Dataset\")\n",
"profile.to_notebook_iframe()"
]
@@ -784,6 +1115,20 @@
"\n",
"Another good package for automated EDA is [dataprep](https://dataprep.ai/)."
]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "f2810c9e",
+ "metadata": {},
+ "source": [
+ "## Summary\n",
+ "\n",
+ "In this chapter, you've learned a variety of tools to help you understand the variation within your data.\n",
+ "You've seen techniques that work with a single variable at a time and with a pair of variables.\n",
+ "This might seem painfully restrictive if you have tens or hundreds of variables in your data, but they're foundation upon which all other techniques are built.\n",
+ "\n",
+ "In the next chapter, we'll focus on the tools we can use to communicate our results."
+ ]
}
],
"metadata": {
@@ -811,7 +1156,7 @@
"name": "python",
"nbconvert_exporter": "python",
"pygments_lexer": "ipython3",
- "version": "3.9.12"
+ "version": "3.10.12"
},
"toc-showtags": true
},
diff --git a/introduction.ipynb b/introduction.ipynb
index e89258f..9cfe8cb 100644
--- a/introduction.ipynb
+++ b/introduction.ipynb
@@ -122,29 +122,17 @@
"\n",
"### R, Julia, and friends\n",
"\n",
- "In this book, you won't learn anything about R, Julia, or any other programming language useful for data science. This isn't because we think these tools are bad. They're not! And in practice, most data science teams use a mix of languages, typically R and Python. However, you may find it easier to learn one tool at a time.\n",
+ "In this book, you won't learn anything about R, Julia, or any other programming language useful for data science. This isn't because we think these tools are bad. They're not! And in practice, most data science teams use a mix of languages. However, you may find it easier to learn one set of tools at a time. In this book you'll see what we think of as the three critical tools for data science:\n",
"\n",
- "This book uses Python, which is usually ranked as the first or second most popular programming language in the world and, just as importantly, it’s also one of the easiest to learn. It’s a general purpose language, which means it can perform a wide range of tasks. This combination of features is why people say Python has a low floor and a high ceiling. It’s also very versatile; the joke goes that Python is the 2nd best language at everything, and there’s some truth to that (although Python is 1st best at some tasks, like machine learning). But a language that covers such a lot of ground is also very useful; and Python is widely used across industry, academia, and the public sector, and is often taught in schools too.\n",
+ "- Python\n",
+ "- SQL\n",
+ "- command line scripting\n",
"\n",
- "We think Python is a great place to start your data science journey because it is the most popular tool for data science and programming more generally, with a large community behind it.\n",
- "\n",
- "## Prerequisites\n",
- "\n",
- "We've made a few assumptions about what you already know in order to get the most out of this book. You should be generally numerically literate, and it's helpful if you have some programming experience already.\n",
- "\n",
- "There are three things you need to run the code in this book: an installation of Python (the programming language), an installation of Visual Studio Code (which runs your Python code), and some key Python 'packages', add-ons to the language that provide extra functionality.\n",
- "\n",
- "An alternative to installing Python on your own computer is to use [Github Codespaces](https://github.com/features/codespaces), a cloud-based version of Visual Studio Code. There is a generous free tier with 60 hours per month of use.\n",
- "\n",
- "### Installing the Python programming language\n",
+ "This book predominantly uses Python, which is usually ranked as the first or second most popular programming language in the world and, just as importantly, it’s also one of the easiest to learn. It’s a general purpose language, which means it can perform a wide range of tasks. This combination of features is why people say Python has a low floor and a high ceiling. It’s also very versatile; the joke goes that Python is the 2nd best language at everything, and there’s some truth to that (although Python is 1st best at some tasks, like machine learning). But a language that covers such a lot of ground is also very useful; and Python is widely used across industry, academia, and the public sector, and is often taught in schools too.\n",
"\n",
- "To download and install the Python language, we'll use the Anaconda distribution of Python, which is available on all major operating systems. To install it, follow the instructions below or watch this video on *[how to install Python using the Anaconda distribution of Python](https://www.youtube.com/watch?v=ZWQwGR5ppnk)*.\n",
- "\n",
- "\n",
- "\n",
- "Download the individual edition of the [Anaconda distribution](https://www.anaconda.com/) of Python for your operating system and install it (on Anaconda's website, this is currently found under Products -> Individual Edition). This will provide you with a Python installation and a host of the most useful libraries. If you get stuck, there are more detailed instructions available for installing the Anaconda distribution of Python [on Windows](https://docs.anaconda.com/anaconda/install/windows/), [on Mac](https://docs.anaconda.com/anaconda/install/mac-os/), and [on Linux](https://docs.anaconda.com/anaconda/install/linux/).\n",
+ "We think Python is a great place to start your data science journey because it is the most popular tool for data science and programming more generally, with a large community behind it.\n",
"\n",
- "You can confirm that you've set up Anaconda correctly by following the [verify installation instructions](https://docs.anaconda.com/anaconda/install/verify-install/) on the Anaconda website. If you're using Windows, you can check if Anaconda has installed properly by opening the 'Anaconda prompt' (a special text-based way to issue commands to your computer) and type `where python`. You should see a path rendered as text in the prompt that includes \"Anaconda3\", for example something like `C:\\Users\\\\Anaconda3\\...`. On Mac and Linux you may need to run `conda init` on your command line to activate your Anaconda Python environment (Mac and Linux usually come with, typically, old versions of Python pre-installed). You can check you've got the right Python with `which python`, which should result in a message back saying `/Users//opt/anaconda3/bin/python`. \n",
+ "## Details about this book\n",
"\n",
"This book was compiled with the following version of Python:"
]
@@ -166,174 +154,6 @@
"print(\"Compiled with Python version:\", sys.version)"
]
},
- {
- "attachments": {},
- "cell_type": "markdown",
- "id": "6e8ffb03",
- "metadata": {},
- "source": [
- "### Installing Visual Studio Code to run Python\n",
- "\n",
- "[Visual Studio Code](https://code.visualstudio.com/) will allow you to run code through Python once you have Python installed on your system. Visual Studio Code is an *integrated development environment*. An integrated development environment is a software application that provides a few tools to make coding easier. The most important of these is a way to write the code itself! IDEs are not the only way to programme, but they are perhaps the most useful. Note that the language and the place you write the language (the IDE) are separate things: the language is a way of processing your instructions, the IDE is where you write those instructions.\n",
- "\n",
- "Here are some of the useful features an IDE might have:\n",
- "\n",
- "- a way to run your code interactively (line-by-line) or all at once\n",
- "\n",
- "- a way to debug (look for errors) in your code\n",
- "\n",
- "- a quick way to access helpful information about commonly used software packages\n",
- "\n",
- "- automatic code formatting, so that your code follows best practice guidelines\n",
- "\n",
- "- auto-completion of your code\n",
- "\n",
- "- automatic code checking for basic errors\n",
- "\n",
- "- colouring your brackets in pairs so you can keep track of the logical order of execution of your code!\n",
- "\n",
- "[Visual Studio Code](https://code.visualstudio.com/) is a free and open source IDE from Microsoft that is available on all major operating systems. Just like Python itself, Visual Studio can be extended with packages, and it is those packages, called extensions in this case, that make it so useful. As well as Python, Visual Studio Code supports a ton of other languages.\n",
- "\n",
- "Download and install Visual Studio Code. If you need some help, the video below will walk you through downloading and installing Visual Studio Code, and then using it to run Python code in both scripts and in notebooks. We'll go through these instructions in detail in the rest of this chapter. As an alternative to the instructions or video below, Microsoft also has a [very short tutorial](https://code.visualstudio.com/docs/python/python-tutorial) on setting it up (ignore the bits about debugging and installing packages for now).\n",
- "\n",
- "\n",
- "\n",
- "*[How to install Visual Studio Code and use it to run Python code](https://www.youtube.com/watch?v=1kKTYsQdaPw)*\n",
- "\n",
- "Once you have Visual Studio Code installed and opened, navigate to the 'extensions' tab on the left hand side vertical bar of icons (it's the one that looks like 4 squares). You'll need to install the [Python extension for VS Code](https://marketplace.visualstudio.com/items?itemName=ms-python.python), which you can search for by using the text box within VS Code's extensions panel.\n",
- "\n",
- "\n",
- "\n",
- "The figure above shows the typical layout of Visual Studio Code. The long vertical panel on the far left-hand side changes what is seen in panels 1 and 2; it currently has the file explorer selected. Let's run through the numbered parts of the figure.\n",
- "\n",
- "1. When the explorer option is selected from the icons to the left of 1 and 2, the contents of the folder that's currently open are shown in 1.\n",
- "2. This is an outline of the key parts of the file that is open in 3.\n",
- "3. This is just a fancy text editor. In the figure above, it's showing a Python script (a file that contains code and has a name that ends in `.py`). Selecting code and pressing Shift + Enter ('Enter' is labelled as 'Return' on some keyboards) will execute that code in 5.\n",
- "4. This is the command line, a place where you can type in commands that your computer will then execute. If you want to try a command, type `date` (Mac/Linux) or `date /t` (Windows).\n",
- "5. This is the interactive Python window, which is where code and code outputs appear after you select and execute them from a script (see 3). It shows the code that you executed and any outputs from that execution—in the screenshot shown, the code has created a plot. The name and version of Python you're using appear at the top of the interactive window.\n",
- "\n",
- "Note that there is lots of useful information arrayed right at the bottom of the window in the blue bar, including the version of Python currently being used by VS Code."
- ]
- },
- {
- "attachments": {},
- "cell_type": "markdown",
- "id": "d9c1931d",
- "metadata": {},
- "source": [
- "### Packages\n",
- "\n",
- "You'll also need to install some Python packages. A Python **package** is a collection of functions, data, and documentation that extends the capabilities of an installed version of Python. Using packages is key to most data science.\n",
- "\n",
- "The Anaconda distribution actually comes with a host of the most useful packages for data science, including **pandas** for data manipulation and **matplotlib** for data visualisation.\n",
- "\n",
- "You can install additional packages from the Visual Studio Code command line (labelled as number 4 in the figure above). To install an extra package, run `pip install **packagename**` in the command line. You could try installing the exploratory data analysis package **skimpy** this way, by running `pip install skimpy`. There'll be more on installing and using packages in {ref}`workflow-packages-and-environments`.\n",
- "\n",
- "If you have problems installing, make sure that you are connected to the internet, and that [PyPI](https://pypi.org/) (the Python package index) isn't blocked by your firewall or proxy."
- ]
- },
- {
- "attachments": {},
- "cell_type": "markdown",
- "id": "9a885521",
- "metadata": {},
- "source": [
- "## Running Python code\n",
- "\n",
- "Now you will create and run your first code. If you get stuck, there's a more in-depth tutorial over at the [VS Code documentation](https://code.visualstudio.com/docs/python/python-tutorial).\n",
- "\n",
- "Create a new folder for your work (perhaps named 'python4DS', no white space), open that folder with Visual Studio Code and create a new file, naming it `hello_world.py`. The file extension, `.py`, is very important as it implicitly tells Visual Studio Code that this is a Python script. In the Visual Studio Code editor, add a single line to the file:\n",
- "\n",
- "```python\n",
- "print('Hello World!')\n",
- "```\n",
- "\n",
- "Save the file.\n",
- "\n",
- "If you named this file with the extension `.py` then VS Code will recognise that it is Python code and you should see the name and version of Python pop up in the blue bar at the bottom of your VS Code window. Make sure that the version of Python displayed here is the Anaconda version that you just installed rather than one that comes built-in with your operating system (this is particularly an issue on Mac). If you have a fresh install of Anaconda's distribution of Python, you'll probably see something like `Python 3.9 64-bit ('base': conda)`. To change which Python version your code uses, click on the version shown in the blue bar and select the version you want. If you've just changed Python version, it can be a good idea to restart VS Code so that all the versions of Python on your system are picked up by it.\n",
- "\n",
- "When you press save, you may get messages about installing extra packages or making Pylance your default language server; just go with VS Code's suggestions here, except the one about the terminal and conda, which you can say no to.\n",
- "\n",
- "Alright, shall we actually run some code? Select/highlight the `print('Hello world!')` text you typed in the file and right-click to bring up some options including 'Run Selection/Line in Terminal' and `Run Selection/Line in Interactive Window'. Because VS Code is a richly featured IDE, there are lots of options for how to run the file. Let's try both of the main ways: via the interactive window and using the \"terminal\" (more on what that is later).\n",
- "\n",
- "The interactive window is a convenient and flexible way to run code that you have open in a script or that you type directly into the interactive window code box. The interactive window will 'remember' any variables that have been assigned (for examples, code statements like `x = 5`), whether they came from running some lines in your script or from you typing them in directly. Working with the interactive window will feel familiar to anyone who has used Stata, Matlab, or R, and is much more suited to the way economists tend to work because it doesn't require you to write the whole script, start to finish, ahead of time. Instead, you can jam, changing code as you go, (re-)running it line by line.\n",
- "\n",
- "To run the code in an interactive window, **right-click and select 'Run Selection/Line in Interactive Window'**. This should cause a new 'interactive' panel to appear within Visual Studio Code, and only the selected line will execute within it. At this point, you may see a message about Visual Studio Code's default behaviour when you press Shift + Enter; for this book, it's good to have Shift + Enter default to running a line in the interactive window. The box below has instructions for how to ensure this always happens.\n",
- "\n",
- "```{admonition} Make code run in the interactive window by default\n",
- ":class: dropdown\n",
- "\n",
- "Open up Visual Studio Code and go to settings (click on the cog in the bottom left-hand corner, then click settings).\n",
- "\n",
- "Type 'python send' into the search box. Depending on your configuration and Visual Studio Code version, you will either see 'Python › Data Science: Send Selection To Interactive Window' or 'Jupyter: Send Selection To Interactive Window'. Make sure that there is a tick in the box.\n",
- "\n",
- "This will ensure that when you hit shift+enter on code scripts, it will execute your code in Visual Studio's interactive window (starting a new window if necessary).\n",
- "```\n",
- "\n",
- "Let's make more use of the interactive window. At the bottom of it, there is a box that says 'Type code here and press shift-enter to run'. Go ahead and type `print('Hello World!')` directly in there to achieve the same effect as running the line from your script. Also, any variables you run in the interactive window (from your script or directly by entering them in the box) will persist.\n",
- "\n",
- "To see how variables persist, type `hello_string = 'Hello World!'` into the interactive window's code entry box and hit shift-enter. If you now type `hello_string` and hit shift+enter, you will see the contents of the variable you just created. You can also click the grid symbol at the top of the interactive window (between the stop symbol and the save file symbol); this is the variable explorer and will pop open a panel showing all of the variables you've created in this interactive session. You should see one called `hello_string` of type `str` with a value `Hello World!`.\n",
- "\n",
- "This shows the two ways of working with the interactive window--running (segments) from a script, or writing code directly in the entry box.\n",
- "\n",
- "```{admonition} Start interactive windows and terminals within your project directory\n",
- ":class: dropdown\n",
- "In Visual Studio Code, you can ensure that the interactive window starts in the root directory of your project by setting \"Jupyter: Notebook File Root\" to `${workspaceFolder}` in the Settings menu. For the integrated command line, change \"Terminal › Integrated: Cwd\" to `${workspaceFolder}` too.\n",
- "```\n",
- "\n",
- "To run code the other way, in the terminal, right-click and select 'Run Python file in terminal'. This will bring up a new panel (called a terminal) *within* Visual Studio Code that runs your entire script from top to bottom-and you should see 'Hello World!' pop up! Although we're trying out running code in the terminal, the typical economics workflow would be to work with the interactive window.\n",
- "\n",
- "```{admonition} Exercise\n",
- "Create a new script that, when run, prints \"Welcome to Python for Data Science\" and run it in an interactive window.\n",
- "```\n"
- ]
- },
- {
- "attachments": {},
- "cell_type": "markdown",
- "id": "a920625f",
- "metadata": {},
- "source": [
- "## Getting help and learning more\n",
- "\n",
- "This book is not an island; there is no single resource that will allow you to master Python. As you start to apply the techniques described in this book to your own data you will soon find questions that we do not answer. This section describes a few tips on how to get help, and to help you keep learning.\n",
- "\n",
- "Some other resources for learning are:\n",
- "\n",
- "- [The Python Data Science Handbook](https://jakevdp.github.io/PythonDataScienceHandbook/)\n",
- "- [Real Python](https://realpython.com/), which has excellent short tutorials that cover Python more broadly (not just data science)\n",
- "- [freeCodeCamp's Python courses](https://www.freecodecamp.org/news/search?query=data%20science%20python), though take care to select one that's at the right level for you\n",
- "- [Coding for Economists](https://aeturrell.github.io/coding-for-economists), which has similar content to this book but is more in depth and aimed at analysts (particularly in economics)\n",
- "\n",
- "If you get stuck with a particular bit of code, start with Google. Typically adding “Python” to a query is enough to restrict it to relevant results. Google is particularly useful for error messages. If you get an error message and you have no idea what it means, try googling it! Chances are that someone else has been confused by it in the past, and there will be help somewhere on the web.\n",
- "\n",
- "If Google doesn’t help, try [stackoverflow](https://stackoverflow.com/). Start by spending a little time searching for an existing answer, including [Python] to restrict your search to questions and answers that use Python. If you don’t find anything useful, prepare a minimal reproducible example or \"reprex\". A good reprex makes it easier for other people to help you, and often you’ll figure out the problem yourself in the course of making it.\n",
- "\n",
- "There are three things you need to include to make your example reproducible: required packages, data, and code.\n",
- "\n",
- "Packages should be loaded at the top of the script, so it’s easy to see which ones the example needs. This is a good time to check that you’re using the latest version of each package; it’s possible you’ve discovered a bug that’s been fixed since you installed the package.\n",
- "\n",
- "You should also spend some time preparing yourself to solve problems before they occur. Investing a little time in learning Python each day will pay off handsomely in the long run.\n",
- "\n",
- "To keep up with the Python data science community more broadly, we recommend following the (#pydata) and (#python) hashtags on Twitter."
- ]
- },
- {
- "attachments": {},
- "cell_type": "markdown",
- "id": "4f2457a2",
- "metadata": {},
- "source": [
- "## How to use this book\n",
- "\n",
- "As well as following this book using your own computer, you can run the code online through a few different options. The first is the easiest to get started with.\n",
- "\n",
- "1. [Google Colab notebooks](https://research.google.com/colaboratory/). You can launch most pages in this book interactively by using the 'Colab' button under the rocket symbol at the top of the page. It will be in the form of a notebook (which mixes code and text) rather than a script (.py file) but the code you write is the same.\n",
- "2. [Github Codespaces](https://github.com/features/codespaces) offer an online version of Visual Studio Code, with a generous free tier of 60 hours per month.\n",
- "3. [Pyolite Notebook](https://jupyterlite.readthedocs.io/en/latest/try/lab). This is a slimmed down version of a Python notebook that runs entirely in your web browser! It doesn't include all of Python's packages but there are enough of the most popular packages for data science to make it useful.\n",
- "4. [Gitpod Workspace](https://www.gitpod.io/). This is a remote, cloud-based version of Visual Studio Code with Python installed and will run Python scripts. Note that the free tier only covers 50 hours per month."
- ]
- },
{
"attachments": {},
"cell_type": "markdown",
diff --git a/prerequisites.ipynb b/prerequisites.ipynb
new file mode 100644
index 0000000..90c78fa
--- /dev/null
+++ b/prerequisites.ipynb
@@ -0,0 +1,287 @@
+{
+ "cells": [
+ {
+ "attachments": {},
+ "cell_type": "markdown",
+ "id": "95f0a171",
+ "metadata": {},
+ "source": [
+ "(prereq)=\n",
+ "# Prerequisites\n",
+ "\n",
+ "Congratulations on starting your data science journey! In this chapter, we're going to help you install or access the tools you need to start learning and doing data science. We've made a few assumptions about what you already know in order to get the most out of this book. You should be generally numerically literate, and it's helpful if you have some programming experience already.\n",
+ "\n",
+ "## Introduction\n",
+ "\n",
+ "We'll be helping you get set up with:\n",
+ "\n",
+ "- an *integrated development environment*, or IDE, a place to write and run code\n",
+ "- an installation of *Python*, so that your computer can interpret and execute Python code\n",
+ "- installing *packages*, which extend the functionality of Python\n",
+ "\n",
+ "While there will be much more information on the how to come, let's first spend a moment explaining *what* these are.\n",
+ "\n",
+ "### Integrated development environment, or IDE\n",
+ "\n",
+ "An *integrated development environment* (IDE) is a software application that provides a few tools to make coding easier. The most important of these is a way to write the code itself! IDEs are not the only way to programme, but they are perhaps the most useful. Note that the language and the place you write the language (the IDE) are separate things: the language is a way of processing your instructions, the IDE is where you write those instructions. There are a lot of integrated development environments (IDEs) out there. This book strongly recommends Microsoft's *Visual Studio Code*, which works on all major operating systems and is one of the most popular. Here are some of the useful features that *Visual Studio Code* provides:\n",
+ "\n",
+ "- a way to run your code interactively (line-by-line) or all at once\n",
+ "\n",
+ "- a way to debug (look for errors) in your code\n",
+ "\n",
+ "- a quick way to access helpful information about commonly used software packages\n",
+ "\n",
+ "- automatic code formatting, so that your code follows best practice guidelines\n",
+ "\n",
+ "- auto-completion of your code when you hit TAB\n",
+ "\n",
+ "- automatic code checking for basic errors\n",
+ "\n",
+ "- colouring your brackets in pairs so you can keep track of the logical order of execution of your code!\n",
+ "\n",
+ "### A Python interpreter\n",
+ "\n",
+ "Python is both a programming language that you can read, and a language that computers can read, interpret, and then carry out instructions based on. For your computer to be able to read and execute Python code, you will need to get Python installed on your computer. There are lots of ways to install a Python \"interpreter\" on your computer, this book recommends the *Anaconda distribution* of Python for its flexibility and simplicity.\n",
+ "\n",
+ "### Packages\n",
+ "\n",
+ "A Python **package** is a collection of functions, data, and documentation that extends the capabilities of an installed version of Python. Using packages is key to most data science because most of the functionality we'll need comes from extra packages. You'll see statetments like `import numpy as np` at the start of many Python code scripts—these are instructions to use an installed package (here one called `numpy`) and to give it a shortened name (`np`, for convenience) in the rest of the script. The functions in the `numpy` package are then accessed through syntax like `np.`; for example, you can take logs with `np.log(x)` where `x` is a variable containing a number. You need only install packages once. \n",
+ "\n",
+ "### Typical workflow\n",
+ "\n",
+ "The typical workflow for analysis with code might be something like this:\n",
+ "\n",
+ "- Open up your *integrated development environment* (IDE)\n",
+ "- Write some code in a script (a text file with code in) in your *IDE*\n",
+ "- If necessary for the analysis that you're doing, install any extra *packages*\n",
+ "- Use the *IDE* to send bits of code from the script, or the entire script, to be executed by *Python* and add-on *packages*, and to display results\n",
+ "\n",
+ "We'll see two ways to achieve this workflow:\n",
+ "\n",
+ "1. Installing an *IDE*, *Python*, and any extra *packages* on your own computer\n",
+ "2. Using a computer in the cloud that you access through your internet browser. The cloud computer has an *IDE* and *Python* built-in, and you can easily install extra *packages* in it too. However, the free version is limited to 60 hours / month.\n",
+ "\n",
+ "You should pick whichever you're more comfortable with! Eventually, you'll probably try both."
+ ]
+ },
+ {
+ "attachments": {},
+ "cell_type": "markdown",
+ "id": "aabdbc7d",
+ "metadata": {},
+ "source": [
+ "## How to get started on your own computer\n",
+ "\n",
+ "### Installing Python\n",
+ "\n",
+ "To download and install Python, we'll use the Anaconda \"distribution\" of Python, which is available on all major operating systems. To install it, follow the instructions below or watch this video on *[how to install Python using the Anaconda distribution of Python](https://www.youtube.com/watch?v=ZWQwGR5ppnk)*.\n",
+ "\n",
+ "\n",
+ "\n",
+ "Download the individual edition of the [Anaconda distribution](https://www.anaconda.com/) of Python for your operating system and install it. This will provide you with a Python installation and a host of the most useful libraries. If you get stuck, there are more detailed instructions available for installing the Anaconda distribution of Python [on Windows](https://docs.anaconda.com/anaconda/install/windows/), [on Mac](https://docs.anaconda.com/anaconda/install/mac-os/), and [on Linux](https://docs.anaconda.com/anaconda/install/linux/).\n",
+ "\n",
+ "You can confirm that you've set up Anaconda correctly by following the [verify installation instructions](https://docs.anaconda.com/free/anaconda/install/verify-install/#conda) on the Anaconda website.\n",
+ "\n",
+ "```{note}\n",
+ "If you're using Windows, you can check if Anaconda has installed properly by opening the 'Anaconda prompt' (a special text-based way to issue commands to your computer) and type `where python`. You should see a path rendered as text in the prompt that includes \"Anaconda3\", for example something like `C:\\Users\\\\Anaconda3\\...`. On Mac and Linux you may need to run `conda init` on your command line to activate your Anaconda Python environment. You can check you've got the right Python with `which python`, which should result in a message back saying `/Users//opt/anaconda3/bin/python`.\n",
+ "```"
+ ]
+ },
+ {
+ "attachments": {},
+ "cell_type": "markdown",
+ "id": "6e8ffb03",
+ "metadata": {},
+ "source": [
+ "### Installing your integrated development environment, Visual Studio Code\n",
+ "\n",
+ "[Visual Studio Code](https://code.visualstudio.com/) is a free and open source IDE from Microsoft that is available on all major operating systems. Just like Python itself, Visual Studio can be extended with packages, and it is those packages, called extensions in this case, that make it so useful. As well as Python, Visual Studio Code supports a ton of other languages.\n",
+ "\n",
+ "Download and install Visual Studio Code. If you need some help, there is a video below that will walk you through downloading and installing Visual Studio Code, and then using it to run Python code in both scripts and in notebooks. We'll go through these instructions in detail in the rest of this chapter.\n",
+ "\n",
+ "\n",
+ "\n",
+ "*[How to install Visual Studio Code and use it to run Python code](https://www.youtube.com/watch?v=1kKTYsQdaPw)*"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "f7b05c4d",
+ "metadata": {},
+ "source": [
+ "## Data science in the cloud\n",
+ "\n",
+ "There are many ways to do data science in the cloud, but we're going to share with you the absolute simplest. For this, you will need to sign up for a [Github Account](https://github.com/). Github is an organisation that's owned by Microsoft and which provides a range of services including a way to back-up code on the cloud, and cloud computing. One of the services offered is *Github Codespaces*. A GitHub Codespace is an online cloud computer that you connect to from your browser window. It has a generous 60 hours free of computing per month.\n",
+ "\n",
+ "```{note}\n",
+ "If you go over the free tier hours on Github Codespaces, your credit card will be charged for any further hours of GitHub Codespaces you use.\n",
+ "```\n",
+ "\n",
+ "Once you've signed up for a Github account, head to [Github Codespaces](https://github.com/codespaces) and click on \"Get Started for Free\". You should see a menu of \"quick start templates\". Under where it says \"Jupyter Notebook\", hit \"Use this template\".\n",
+ "\n",
+ "You will find that a new page loads with several panels in. This is an online version of Visual Studio Code that works much like if you had installed it on your own computer. It will already have a version of Python installed—you can check which one by running `python --version` in the terminal. The terminal is usually found in the lowest panel of Visual Studio Code, and, in Codespaces, will typically display a welcome message."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "b474cbdd",
+ "metadata": {},
+ "source": [
+ "## Running your first Python code\n",
+ "\n",
+ "### Getting to grips with Visual Studio Code\n",
+ "\n",
+ "Once you have Visual Studio Code installed and opened (either on your own computer or in the cloud), navigate to the 'extensions' tab on the left hand side vertical bar of icons (it's the one that looks like 4 squares). You'll need to install the *Python extension for VS Code*, which you can search for by using the text box within VS Code's extensions panel. If you're using the cloud version, you may find that it's already installed.\n",
+ "\n",
+ "There are some other extensions it's useful to have and install (if they aren't already):\n",
+ "\n",
+ "- Jupyter\n",
+ "- Pylance\n",
+ "- indent-rainbow\n",
+ "\n",
+ "Although you won't have any Python code to play with yet, or an interactive window to execute that Python code, it's worth us spending a brief moment familiarising ourselves with the different bits of a *typical* view in Visual Studio Code.\n",
+ "\n",
+ "\n",
+ "\n",
+ "The figure above shows the typical layout of Visual Studio Code once you have a Python session running, and a Python script open. The long vertical panel on the far left-hand side changes what is seen in panels 1 and 2; it currently has the file explorer selected. Let's run through the numbered parts of the figure.\n",
+ "\n",
+ "1. When the explorer option is selected from the icons to the left of 1 and 2, the contents of the folder that's currently open are shown in 1.\n",
+ "2. This is an outline of the key parts of the file that is open in 3.\n",
+ "3. This is just a fancy text editor. In the figure above, it's showing a Python script (a file that contains code and has a name that ends in `.py`). Shortly, we'll see how selecting code and pressing Shift + Enter ('Enter' is labelled as 'Return' on some keyboards) will execute code whose results appear in panel 5.\n",
+ "4. This is the command line or *terminal*, a place where you can type in commands that your computer will then execute. If you want to try a command, type `date` (Mac/Linux) or `date /t` (Windows). This is where we install extra *packages*.\n",
+ "5. This is the interactive Python window, which is where code and code outputs appear after you select and execute them from a script (see 3). It shows the code that you executed and any outputs from that execution—in the screenshot shown, the code has created a plot. The name and version of Python you're using appear at the top of the interactive window.\n",
+ "\n",
+ "Note that there is lots of useful information arrayed right at the bottom of the window in the blue bar, including the version of Python currently being used by VS Code."
+ ]
+ },
+ {
+ "attachments": {},
+ "cell_type": "markdown",
+ "id": "9a885521",
+ "metadata": {},
+ "source": [
+ "### Running Python code\n",
+ "\n",
+ "Now you will create and run your first code. If you get stuck, there's a more in-depth tutorial over at the [VS Code documentation](https://code.visualstudio.com/docs/python/python-tutorial).\n",
+ "\n",
+ "In Visual Studio Code, click on the \"Explorer\" symbol (some files on the left-hand side of the screen) to bring up a file explorer. Check you're in a good location on your computer to try things out and, if not, change the folder you're in using File -> Open Folder until you're happy.\n",
+ "\n",
+ "Now, still with the explorer panel open, click on the symbol that looks like a blank piece of paper with a \"+\" sign on it. This will create a new file, and your cursor should move to name it. Name it `hello_world.py`. The file extension, `.py`, is very important as it implicitly tells Visual Studio Code that this is a Python script.\n",
+ "\n",
+ "In the Visual Studio Code editor, add a single line to the file:\n",
+ "\n",
+ "```python\n",
+ "print('Hello World!')\n",
+ "```\n",
+ "\n",
+ "Save the file.\n",
+ "\n",
+ "If you named this file with the extension `.py` then VS Code will recognise that it is Python code and you should see the name and version of Python pop up in the bar at the bottom of your VS Code window. (You can have multiple versions of Python installed—if you ever want to change which Python version your code uses, click on the version shown in the bar and select the version you want.)\n",
+ "\n",
+ "Alright, shall we actually **run some code**? Select/highlight the `print(\"Hello world!\")` text you typed in the file and right-click. You'll get a lot of options here, but the one you want is **\"Run Selection/Line in Interactive Window\"**.\n",
+ "\n",
+ "This should cause a new 'interactive' panel to appear within Visual Studio Code, and, hey presto you should see:\n",
+ "\n",
+ "```python\n",
+ "print(\"Hello world!\")\n",
+ "```\n",
+ "```text\n",
+ "Hello world!\n",
+ "```\n",
+ "\n",
+ "The *interactive window* is a convenient and flexible way to run code that you have open in a script or that you type directly into the interactive window code box. The interactive window will 'remember' any variables that have been assigned (for examples, code statements like `x = 5`), whether they came from running some lines in your script or from you typing them in directly. Working with the interactive window will feel familiar to anyone who has used Stata, Matlab, or R. It doesn't require you to write the whole script, start to finish, ahead of time. Instead, you can jam, changing code as you go, (re-)running it line by line.\n",
+ "\n",
+ "It would be cumbersome to have to right-click every time we wanted to run some code, so we're going to make a *keyboard shortcut* to send whatever code is highlighted to the interactive window to be executed. To do this:\n",
+ "\n",
+ "- Open up the Visual Studio Code configuration menu (the cog on the lower left-hand side)\n",
+ "- Go to Settings\n",
+ "- Type \"jupyter send\" in the box to make an entry \"Interactive Window > Text Editor: Execute Selection\" appear\n",
+ "- Ensure the box next to this entry is ticked\n",
+ "\n",
+ "Now return to your script, put your cursor on the line with `print(\"Hello world!\")` on, and hit Shift+Enter. You should see \"Hello world!\" appear again, only this time, it was much easier.\n",
+ "\n",
+ "\n",
+ "```{admonition} Running code in the terminal instead\n",
+ ":class: dropdown\n",
+ "\n",
+ "The interactive window isn't the only way to run code; you can do it in the terminal too. This is less popular for data science, but it does occasionally have its uses. If you want to do this, right-click on the selected code and choose \"Run Python -> Run Selection/Line in Terminal\".\n",
+ "```\n",
+ "\n",
+ "Let's make more use of the *interactive window*. At the bottom of it, there is a box that says 'Type code here and press shift-enter to run'. Go ahead and type `print('Hello World!')` directly in there to achieve the same effect as running the line from your script. Also, any variables you run in the interactive window (from your script or directly by entering them in the box) will persist.\n",
+ "\n",
+ "To see how variables persist, type `hello_string = 'Hello World!'` into the interactive window's code entry box and hit shift-enter. If you now type `hello_string` and hit shift+enter, you will see the contents of the variable you just created. You can also click the grid symbol at the top of the interactive window (between the stop symbol and the save file symbol); this is the variable explorer and will pop open a panel showing all of the variables you've created in this interactive session. You should see one called `hello_string` of type `str` with a value `Hello World!`.\n",
+ "\n",
+ "This shows the two ways of working with the interactive window--running (segments) from a script, or writing code directly in the entry box. It doesn't matter which way you entered variables, they will all be remembered within that session in your interactive window.\n",
+ "\n",
+ "```{admonition} Start interactive windows and terminals within your project directory\n",
+ ":class: dropdown\n",
+ "In Visual Studio Code, you can ensure that the interactive window starts in the root directory of your project by setting \"Jupyter: Notebook File Root\" to `${workspaceFolder}` in the Settings menu. For the integrated command line, change \"Terminal › Integrated: Cwd\" to `${workspaceFolder}` too.\n",
+ "```\n",
+ "\n",
+ "```{admonition} Exercise\n",
+ "Create a new script that, when run, prints \"Welcome to Python for Data Science\" and run it in an interactive window.\n",
+ "```"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "6a191ca0",
+ "metadata": {},
+ "source": [
+ "### Installing Packages\n",
+ "\n",
+ "We use the *terminal* or *command line* within Visual Studio Code to install additional Python packages. In the figure earlier in the Chapter, this is labelled as panel number 4.\n",
+ "\n",
+ "To install an extra package, run `pip install **packagename**` in the command line. You could try installing the data analysispackage **polars** this way, by running `pip install polars`. We'll see how to use **polars** in later chapters, but if you want to know if it installed correctly, just look for the message saying \"Successfully installed polars\" followed by the version number.\n",
+ "\n",
+ "There'll be more on installing and using packages in {ref}`workflow-packages-and-environments`.\n",
+ "\n",
+ "If you have problems installing, make sure that you are connected to the internet, and that [PyPI](https://pypi.org/) (the Python package index) isn't blocked by your firewall or proxy."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "ba117896",
+ "metadata": {},
+ "source": [
+ "## Alternative ways to run the code from the book\n",
+ "\n",
+ "As well as following this book using your own computer or on the cloud via Github Codespaces, you can run the code online through a few other options. The first is the easiest to get started with.\n",
+ "\n",
+ "1. [Google Colab notebooks](https://research.google.com/colaboratory/). Free for most use. You can launch most pages in this book interactively by using the 'Colab' button under the rocket symbol at the top of the page. It will be in the form of a notebook (which mixes code and text) rather than a script (.py file) but the code you write is the same.\n",
+ "2. [Gitpod Workspace](https://www.gitpod.io/). An alternative to Codespaces. This is a remote, cloud-based version of Visual Studio Code with Python installed and will run Python scripts. Note that the free tier covers 50 hours per month."
+ ]
+ }
+ ],
+ "metadata": {
+ "interpreter": {
+ "hash": "9d7534ecd9fbc7d385378f8400cf4d6cb9c6175408a574f1c99c5269f08771cc"
+ },
+ "jupytext": {
+ "cell_metadata_filter": "-all",
+ "encoding": "# -*- coding: utf-8 -*-",
+ "formats": "md:myst",
+ "main_language": "python"
+ },
+ "kernelspec": {
+ "display_name": "Python 3 (ipykernel)",
+ "language": "python",
+ "name": "python3"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.9.12"
+ },
+ "toc-showtags": true
+ },
+ "nbformat": 4,
+ "nbformat_minor": 5
+}
diff --git a/rectangling.ipynb b/rectangling.ipynb
index fc7af61..bfa2ae9 100644
--- a/rectangling.ipynb
+++ b/rectangling.ipynb
@@ -6,11 +6,11 @@
"metadata": {},
"source": [
"(rectangling)=\n",
- "# Rectangling\n",
+ "# Nested Data\n",
"\n",
"## Introduction\n",
"\n",
- "In this chapter, you'll learn the art of data **rectangling**, taking data that is fundamentally tree-like and converting it into a rectangular data frames made up of rows and columns. This is important because hierarchical data is surprisingly common, especially when working with data that comes from a web API (such as you will see in {ref}`webscraping-and-apis`).\n",
+ "In this chapter, you'll learn about **nested data**, working with data that is fundamentally tree-like and (often) converting it into a rectangular data frames made up of rows and columns. This is important because nested data is surprisingly common, especially when working with data that comes from a web API (such as you will see in {ref}`webscraping-and-apis`).\n",
"\n",
"To learn about rectangling, you'll first learn about lists, dictionaries, and the JSON format, as these are the data structures that are most often used to work with hierarchical data in Python. Then you'll learn about some functions that can help you turn hierarchical data into 'tidy' data in columns and rows. We'll then show you a few case studies, applying these simple function multiple times to solve real complex problems.\n"
]
diff --git a/spreadsheets.ipynb b/spreadsheets.ipynb
index 75d5710..e234fac 100644
--- a/spreadsheets.ipynb
+++ b/spreadsheets.ipynb
@@ -174,7 +174,7 @@
"\n",
"An important feature that distinguishes spreadsheets from flat files is the notion of multiple sheets. The figure below shows an Excel spreadsheet with multiple sheets. The data come from the **palmerpenguins** dataset {cite}`horst2020palmerpenguins`. Each sheet contains information on penguins from a different island where data were collected.\n",
"\n",
- "\n",
+ "\n",
"\n",
"You can read a single sheet using the following command (so as not to show the whole file, we'll use `.head()` to just show the first 5 rows):"
]
@@ -265,7 +265,7 @@
"\n",
"The figure below shows such a spreadsheet: in the middle of the sheet is what looks like a data frame but there is extraneous text in cells above and below the data.\n",
"\n",
- "\n",
+ "\n",
"\n"
]
},
@@ -383,7 +383,7 @@
"source": [
"The figure below shows what the data looks like in Excel.\n",
"\n",
- ""
+ ""
]
},
{
diff --git a/vis-layers.ipynb b/vis-layers.ipynb
new file mode 100644
index 0000000..038b5c3
--- /dev/null
+++ b/vis-layers.ipynb
@@ -0,0 +1,1006 @@
+{
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "id": "95f0a171",
+ "metadata": {},
+ "source": [
+ "(vis-layers)=\n",
+ "# Layers\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "9fdd3b8a",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "\n",
+ "In the previous chapters, you've learned much more than just how to make scatterplots, bar charts, and boxplots.\n",
+ "You learned a foundation that you can use to make *any* type of plot with **lets-plot**.\n",
+ "\n",
+ "In this chapter, you'll expand on that foundation as you learn about the layered grammar of graphics.\n",
+ "We'll start with a deeper dive into aesthetic mappings, geometric objects, and facets.\n",
+ "Then, you will learn about statistical transformations **lets-plot** makes under the hood when creating a plot.\n",
+ "These transformations are used to calculate new values to plot, such as the heights of bars in a bar plot or medians in a box plot.\n",
+ "You will also learn about position adjustments, which modify how geoms are displayed in your plots.\n",
+ "Finally, we'll briefly introduce coordinate systems.\n",
+ "\n",
+ "We will not cover every single function and option for each of these layers, but we will walk you through the most important and commonly used functionality provided by **lets-plot**."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "17575f3a",
+ "metadata": {},
+ "source": [
+ "### Prerequisites\n",
+ "\n",
+ "You will need to install the **letsplot** package for this chapter, as well as **pandas**.\n",
+ "\n",
+ "In your Python session, import the libraries we'll be using:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "a86fb211",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import pandas as pd\n",
+ "from lets_plot import *\n",
+ "from lets_plot.geo_data import *\n",
+ "from lets_plot.mapping import as_discrete\n",
+ "\n",
+ "LetsPlot.setup_html()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "55b00fde",
+ "metadata": {},
+ "source": [
+ "## Aesthetic mappings\n",
+ "\n",
+ "> \"The greatest value of a picture is when it forces us to notice what we never expected to see.\" --- John Tukey\n",
+ "\n",
+ "We're going to use the `mpg` dataset for this section, so let's download it."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "39a6d993",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "mpg = pd.read_csv(\n",
+ " \"https://vincentarelbundock.github.io/Rdatasets/csv/ggplot2/mpg.csv\", index_col=0\n",
+ ")\n",
+ "\n",
+ "mpg = mpg.astype(\n",
+ " {\n",
+ " \"manufacturer\": \"category\",\n",
+ " \"model\": \"category\",\n",
+ " \"displ\": \"double\",\n",
+ " \"year\": \"int64\",\n",
+ " \"cyl\": \"int64\",\n",
+ " \"trans\": \"category\",\n",
+ " \"drv\": \"category\",\n",
+ " \"cty\": \"double\",\n",
+ " \"hwy\": \"double\",\n",
+ " \"fl\": \"category\",\n",
+ " \"class\": \"category\",\n",
+ " }\n",
+ ")\n",
+ "mpg.head()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "6d6f1307",
+ "metadata": {},
+ "source": [
+ "Among the variables in `mpg` are:\n",
+ "\n",
+ "1. `displ`: A car's engine size, in liters.\n",
+ " A numerical variable.\n",
+ "\n",
+ "2. `hwy`: A car's fuel efficiency on the highway, in miles per gallon (mpg).\n",
+ " A car with a low fuel efficiency consumes more fuel than a car with a high fuel efficiency when they travel the same distance.\n",
+ " A numerical variable.\n",
+ "\n",
+ "3. `class`: Type of car.\n",
+ " A categorical variable.\n",
+ "\n",
+ "Let's start by visualising the relationship between `displ` and `hwy` for various `class`es of cars.\n",
+ "We can do this with a scatterplot where the numerical variables are mapped to the `x` and `y` aesthetics and the categorical variable is mapped to an aesthetic like `color` or `shape`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "fe77349a",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"displ\", y=\"hwy\", color=\"class\")) + geom_point())"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "e77b5640",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"displ\", y=\"hwy\", shape=\"class\")) + geom_point())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "53e51510",
+ "metadata": {},
+ "source": [
+ "Similarly, we can map `class` to `size` or `alpha` aesthetics as well, which control the shape and the transparency of the points, respectively."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "ef221330",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"displ\", y=\"hwy\", size=\"class\")) + geom_point())"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "d042255e",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"displ\", y=\"hwy\", alpha=\"class\")) + geom_point())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "ec07349f",
+ "metadata": {},
+ "source": [
+ "While we are able to do it, mapping an unordered discrete (categorical) variable (`class`) to an ordered aesthetic variable (`size` or `alpha`) is generally not a good idea because it implies a ranking that does not in fact exist.\n",
+ "\n",
+ "Once you map an aesthetic, **lets-plot** takes care of the rest.\n",
+ "It selects a reasonable scale to use with the aesthetic, and it constructs a legend that explains the mapping between levels and values.\n",
+ "For x and y aesthetics, **lets-plot** does not create a legend, but it creates an axis line with tick marks and a label.\n",
+ "The axis line provides the same information as a legend; it explains the mapping between locations and values.\n",
+ "\n",
+ "You can also set the visual properties of your geom manually as an argument of your geom function (*outside* of `aes()`) instead of relying on a variable mapping to determine the appearance.\n",
+ "For example, we can make all of the points in our plot blue:\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "618edcb4",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"displ\", y=\"hwy\")) + geom_point(color=\"blue\"))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "52611640",
+ "metadata": {},
+ "source": [
+ "Here, the colour doesn't convey information about a variable, but only changes the appearance of the plot.\n",
+ "You'll need to pick a value that makes sense for that aesthetic:\n",
+ "\n",
+ "- The name of a color as a character string, e.g., `color = \"blue\"`\n",
+ "- The size of a point in mm, e.g., `size = 1`\n",
+ "- The shape of a point as a number, e.g, `shape = 1`.\n",
+ "\n",
+ "Try changing the above plot but, instead of specifying colour, try specifying the shape aesthetic. What do you get with shape set to 1, 2, or 3?"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "37d49ff8",
+ "metadata": {},
+ "source": [
+ "So far we have discussed aesthetics that we can map or set in a scatterplot, when using a point geom.\n",
+ "\n",
+ "The specific aesthetics you can use for a plot depend on the geom you use to represent the data.\n",
+ "In the next section we dive deeper into geoms."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "3f1da019",
+ "metadata": {},
+ "source": [
+ "1. Create a scatterplot of `hwy` vs. `displ` where the points are pink filled in triangles.\n",
+ "\n",
+ "2. Why does the following code not result in a plot with blue points?\n",
+ "\n",
+ " ```python\n",
+ " (\n",
+ " ggplot(mpg) + \n",
+ " geom_point(aes(x = \"displ\", y = \"hwy\", color = \"blue\"))\n",
+ " )\n",
+ " ```\n",
+ "\n",
+ "3. What does the `stroke` aesthetic do?\n",
+ " What shapes does it work with?\n",
+ " (Hint: use `stroke` in the global aesthetic and `shape` in `geom_point`)\n",
+ "\n",
+ "4. Try changing the last plot from above but, instead of specifying colour, try specifying the shape aesthetic. What do you get with shape set to 1, 2, or 3?"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "83aa98f0",
+ "metadata": {},
+ "source": [
+ "## Geometric objects\n",
+ "\n",
+ "How are these two plots similar?"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "277a4c0f",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"displ\", y=\"hwy\")) + geom_point(size=4))"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "07247ba9",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"displ\", y=\"hwy\")) + geom_smooth(method=\"loess\", size=2))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "58824a10",
+ "metadata": {},
+ "source": [
+ "Both plots contain the same x variable, the same y variable, and both describe the same data.\n",
+ "But the plots are not identical.\n",
+ "Each plot uses a different geometric object, geom, to represent the data.\n",
+ "The plot on the left uses the point geom, and the plot on the right uses the smooth geom, a smooth line fitted to the data.\n",
+ "\n",
+ "To change the geom in your plot, change the geom function that you add to `ggplot()`."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "0db26c6c",
+ "metadata": {},
+ "source": [
+ "Every geom function in **lets-plot** takes a `mapping` argument, either defined locally in the geom layer or globally in the `ggplot()` layer.\n",
+ "However, not every aesthetic works with every geom.\n",
+ "You could set the shape of a point, but you couldn't set the \"shape\" of a line.\n",
+ "If you try, **lets-plot** will silently ignore that aesthetic mapping.\n",
+ "On the other hand, you *could* set the linetype of a line.\n",
+ "`geom_smooth()` will draw a different line, with a different linetype, for each unique value of the variable that you map to linetype.\n",
+ "\n",
+ "Let's take a look:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "4b20c825",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"displ\", y=\"hwy\", line=\"drv\")) + geom_smooth(method=\"loess\"))"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "84df3e78",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"displ\", y=\"hwy\", linetype=\"drv\")) + geom_smooth(method=\"loess\"))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "7b114911",
+ "metadata": {},
+ "source": [
+ "Here, `geom_smooth()` separates the cars into three lines based on their `drv` value, which describes a car's drive train.\n",
+ "One line describes all of the points that have a `4` value, one line describes all of the points that have an `f` value, and one line describes all of the points that have an `r` value.\n",
+ "Here, `4` stands for four-wheel drive, `f` for front-wheel drive, and `r` for rear-wheel drive.\n",
+ "\n",
+ "If this is too confusing, we can make it clearer by overlaying the lines on top of the raw data and then coloring everything according to `drv`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "c9e8d92f",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\", color=\"drv\"))\n",
+ " + geom_point()\n",
+ " + geom_smooth(aes(linetype=\"drv\"), method=\"loess\")\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "b6392da1",
+ "metadata": {},
+ "source": [
+ "Notice that this plot contains two geoms in the same graph.\n",
+ "\n",
+ "Many geoms, like `geom_smooth()`, use a single geometric object to display multiple rows of data.\n",
+ "For these geoms, you can set the `group` aesthetic to a categorical variable to draw multiple objects.\n",
+ "**lets-plot** will draw a separate object for each unique value of the grouping variable.\n",
+ "In practice, **lets-plot** will automatically group the data for these geoms whenever you map an aesthetic to a discrete variable.\n",
+ "It is convenient to rely on this feature because the `group` aesthetic by itself does not add a legend or distinguishing features to the geoms.\n",
+ "\n",
+ "Note that if you place mappings in a geom function, **lets-plot** will treat them as local mappings for the layer.\n",
+ "It will use these mappings to extend or overwrite the global mappings *for that layer only*.\n",
+ "This makes it possible to display different aesthetics in different layers."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "b3916558",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"displ\", y=\"hwy\")) + geom_point(aes(color=\"class\")) + geom_smooth())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "88708546",
+ "metadata": {},
+ "source": [
+ "You can use the same idea to specify different data for each layer.\n",
+ "Here, we use red points as well as open circles to highlight two-seater cars.\n",
+ "The local data argument in `geom_point()` overrides the global data argument in `ggplot()` for that layer only.\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "38870eb5",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\"))\n",
+ " + geom_point()\n",
+ " + geom_point(data=mpg.loc[mpg[\"class\"] == \"2seater\", :], color=\"red\", size=2)\n",
+ " + geom_point(\n",
+ " data=mpg.loc[mpg[\"class\"] == \"2seater\", :], shape=1, size=3, color=\"red\"\n",
+ " )\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "35a3d017",
+ "metadata": {},
+ "source": [
+ "Geoms are the fundamental building blocks of **lets-plot**.\n",
+ "You can completely transform the look of your plot by changing its geom, and different geoms can reveal different features of your data.\n",
+ "\n",
+ "**lets-plot** provides over 40 geoms but these don't cover all possible plots one could make. You can find an overview at the relevant part of the [**lets-plot** documentation](https://lets-plot.org/pages/api.html#geometries).\n",
+ "\n",
+ "If you need a geom that is not included, you have three main options:\n",
+ "1. Look for packages that extend **lets-plot** and that do what you need\n",
+ "2. Raise an issue on the [**lets-plot** Github page](https://github.com/JetBrains/lets-plot) requesting it as a new feature—but bear in mind that it might not be a priority for the maintainers, and there's no guarantee that they'll add it, depending on how useful it is for others and how easy it is to implemet.\n",
+ "3. Turn to an imperative plotting package that gives you fine-grained control so you can build your own chart from the ground up—[**matplotlib**](https://matplotlib.org/) is absolutely excellent for this.\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "39a12f36",
+ "metadata": {},
+ "source": [
+ "### Exercises\n",
+ "\n",
+ "1. What geom would you use to draw a line chart?\n",
+ " A boxplot?\n",
+ " A histogram?\n",
+ " An area chart?\n",
+ "\n",
+ "2. What effect would running the previous example:\n",
+ "\n",
+ " ```python\n",
+ " (\n",
+ " ggplot(mpg, aes(x = \"displ\", y = \"hwy\", alpha = \"class\")) +\n",
+ " geom_point()\n",
+ " )\n",
+ " ```\n",
+ " with the keyword argument `show_legend=False` have on the chart generated by this code?\n",
+ "\n",
+ "3. What does the `se` argument to `geom_smooth()` do?\n",
+ "\n",
+ "4. Recreate the Python code necessary to generate the following graph.\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "ae75c5c1",
+ "metadata": {
+ "tags": [
+ "remove-cell"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "(\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\", color=\"drv\"))\n",
+ " + geom_smooth(aes(group=\"drv\"), se=False, method=\"loess\")\n",
+ " + geom_point()\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "4c1d45ab",
+ "metadata": {},
+ "source": [
+ "## Facets\n",
+ "\n",
+ "In {ref}`data-visualise`, you learned about faceting with `facet_wrap()`, which splits a plot into subplots that each display one subset of the data based on a categorical variable.\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "cb651300",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"displ\", y=\"hwy\")) + geom_point() + facet_wrap(\"cyl\"))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "0cd9ef67",
+ "metadata": {},
+ "source": [
+ "To facet your plot with the combination of two variables, switch from `facet_wrap()` to `facet_grid()`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "61481052",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"displ\", y=\"hwy\")) + geom_point() + facet_grid(\"drv\", \"cyl\"))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "4f502141",
+ "metadata": {},
+ "source": [
+ "By default each of the facets share the same scale and range for x and y axes.\n",
+ "This is useful when you want to compare data across facets, and is the recommended default, but it can be limiting when you want to visualise the relationship within each facet better.\n",
+ "Setting the `scales` argument in a faceting function to `\"free\"` will allow for different axis scales across both rows and columns, `\"free_x\"` will allow for different scales across rows, and `\"free_y\"` will allow for different scales across columns.\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "adcd9079",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(\n",
+ " ggplot(mpg, aes(x=\"displ\", y=\"hwy\"))\n",
+ " + geom_point()\n",
+ " + facet_grid(\"drv\", \"cyl\", scales=\"free_y\")\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "ceb2a354",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg) + geom_point(aes(x=\"displ\", y=\"hwy\")) + facet_wrap(\"class\", nrow=2))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "5e3a949f",
+ "metadata": {},
+ "source": [
+ "### Exercises\n",
+ "\n",
+ "1. What happens if you facet on a continuous variable?\n",
+ "\n",
+ "2. What do the empty cells in plot with `facet_grid(\"drv\", \"cyl\")` mean?\n",
+ " Run the following code.\n",
+ " How do they relate to the resulting plot?\n",
+ "\n",
+ " ```python\n",
+ " (\n",
+ " ggplot(mpg) + \n",
+ " geom_point(aes(x = \"drv\", y = \"cyl\"))\n",
+ " )\n",
+ " ```\n",
+ "\n",
+ "3. What plots does the following code make?\n",
+ " What does omitting the second variable do?\n",
+ "\n",
+ " ```python\n",
+ " (\n",
+ " ggplot(mpg) + \n",
+ " geom_point(aes(x = \"displ\", y = \"hwy\")) +\n",
+ " facet_grid(\"drv\")\n",
+ " )\n",
+ "\n",
+ " (\n",
+ " ggplot(mpg) + \n",
+ " geom_point(aes(x = displ, y = \"hwy\")) +\n",
+ " facet_grid(\"cyl\")\n",
+ " )\n",
+ " ```\n",
+ "\n",
+ "4. Take the first faceted plot in this section:\n",
+ "\n",
+ " ```python\n",
+ " (\n",
+ " ggplot(mpg) + \n",
+ " geom_point(aes(x = \"displ\", y = \"hwy\")) + \n",
+ " facet_wrap(\"class\", nrow = 2)\n",
+ " )\n",
+ " ```\n",
+ "\n",
+ " What are the advantages to using faceting instead of the color aesthetic?\n",
+ " What are the disadvantages?\n",
+ " How might the balance change if you had a larger dataset?\n",
+ "\n",
+ "\n",
+ "5. Read `help(facet_wrap)` or hover your mouse over `facet_wrap` in Visual Studio Code.\n",
+ " What does `nrow` do?\n",
+ " What does `ncol` do?\n",
+ " What other options control the layout of the individual panels?\n",
+ " Why doesn't `facet_grid()` have `nrow` and `ncol` arguments?\n",
+ "\n",
+ "6. Recreate the following plot using `facet_wrap()` instead of `facet_grid()`.\n",
+ " How do the positions of the facet labels change?\n",
+ "\n",
+ " ```python\n",
+ " (\n",
+ " ggplot(mpg) + \n",
+ " geom_point(aes(x = \"displ\", y = \"hwy\")) +\n",
+ " facet_grid(\"drv\")\n",
+ " )\n",
+ " ```"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "cacf1fb5",
+ "metadata": {},
+ "source": [
+ "## Statistical transformations\n",
+ "\n",
+ "Consider a basic bar chart, drawn with `geom_bar()` or `geom_col()`.\n",
+ "The following chart displays the total number of diamonds in the `diamonds` dataset, grouped by `cut`.\n",
+ "The `diamonds` dataset contains information on \\~54,000 diamonds, including the `price`, `carat`, `color`, `clarity`, and `cut` of each diamond. We'll load it in a moment.\n",
+ "The chart shows that more diamonds are available with high quality cuts than with low quality cuts."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "f379e31b",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "diamonds = pd.read_csv(\n",
+ " \"https://vincentarelbundock.github.io/Rdatasets/csv/ggplot2/diamonds.csv\",\n",
+ " index_col=0,\n",
+ ")\n",
+ "diamonds_cut_order = [\"Fair\", \"Good\", \"Very Good\", \"Premium\", \"Ideal\"]\n",
+ "diamonds[\"cut\"] = diamonds[\"cut\"].astype(\n",
+ " pd.CategoricalDtype(categories=diamonds_cut_order, ordered=True)\n",
+ ")\n",
+ "diamonds.head()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "d8faf1ab",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(diamonds, aes(x=\"cut\")) + geom_bar())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "a27666cf",
+ "metadata": {},
+ "source": [
+ "On the x-axis, the chart displays `cut`, a variable from `diamonds`.\n",
+ "On the y-axis, it displays count, but count is not a variable in `diamonds`!\n",
+ "Where does count come from?\n",
+ "Many graphs, like scatterplots, plot the raw values of your dataset.\n",
+ "Other graphs, like bar charts, calculate new values to plot:\n",
+ "\n",
+ "- Bar charts, histograms, and frequency polygons bin your data and then plot bin counts, the number of points that fall in each bin.\n",
+ "\n",
+ "- Smoothers fit a model to your data and then plot predictions from the model.\n",
+ "\n",
+ "- Boxplots compute the five-number summary of the distribution and then display that summary as a specially formatted box.\n",
+ "\n",
+ "The algorithm used to calculate new values for a graph is called a **stat**, short for statistical transformation.\n",
+ "The figure below shows how this process works with `geom_bar()`.\n",
+ "\n",
+ "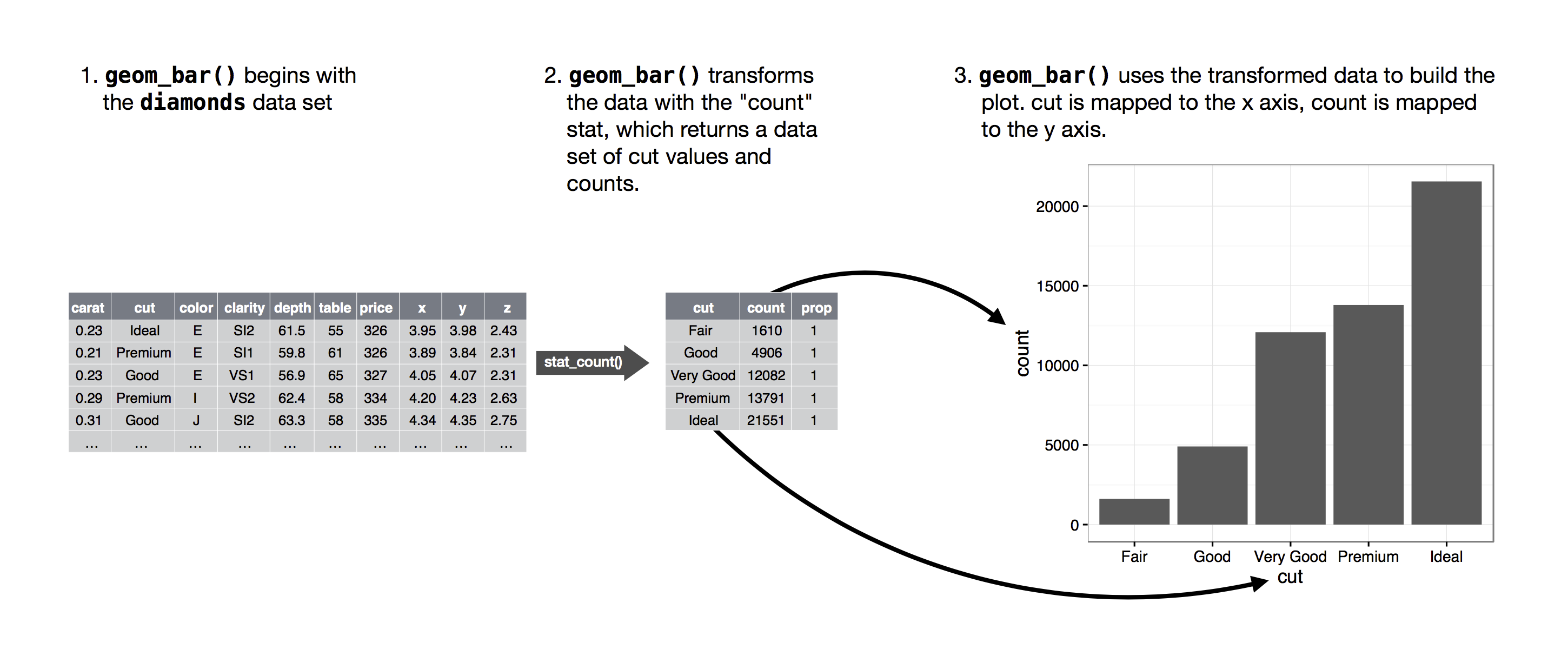\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "62519f73",
+ "metadata": {},
+ "source": [
+ "You can learn which stat a geom uses by inspecting the default value for the `stat` argument.\n",
+ "For example, `help(geom_bar)` (or hovering your mouse over the function written in code) shows that the default value for `stat` is \"count\", which means that `geom_bar()` uses counts of the number of occurrences.\n",
+ "\n",
+ "Every geom has a default stat; and every stat has a default geom.\n",
+ "This means that you can typically use geoms without worrying about the underlying statistical transformation.\n",
+ "However, there are some reasons why you might need to use a stat explicitly; for example, you might want to override the default stat. In the code below, we change the stat of `geom_bar()` from count (the default) to identity. This lets us map the height of the bars to the raw values of a y variable."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "ca772dd5",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(\n",
+ " ggplot(\n",
+ " diamonds.value_counts(\"cut\").reset_index(name=\"counts\"),\n",
+ " aes(x=\"cut\", y=\"counts\"),\n",
+ " )\n",
+ " + geom_bar(stat=\"identity\")\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "e365aaaf",
+ "metadata": {},
+ "source": [
+ "## Position adjustments\n",
+ "\n",
+ "There's one more piece of magic associated with bar charts.\n",
+ "You can colour a bar chart using either the `color` aesthetic, or, more usefully, the `fill` aesthetic:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "f8da7d91",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"drv\", color=\"drv\")) + geom_bar())"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "088e7550",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"drv\", fill=\"drv\")) + geom_bar())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "c3f7a0a3",
+ "metadata": {},
+ "source": [
+ "Note what happens if you map the fill aesthetic to another variable, like `class`: the bars are automatically stacked.\n",
+ "Each colored rectangle represents a combination of `drv` and `class`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "181c70d2",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"drv\", fill=\"class\")) + geom_bar())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "b3f74621",
+ "metadata": {},
+ "source": [
+ "The stacking is performed automatically using the **position adjustment** specified by the `position` argument.\n",
+ "If you don't want a stacked bar chart, you can use one of three other options: `\"identity\"`, `\"dodge\"` or `\"fill\"`.\n",
+ "\n",
+ "- `position = \"identity\"` will place each object exactly where it falls in the context of the graph.\n",
+ " This is not very useful for bars, because it overlaps them.\n",
+ " To see that overlapping we usually need to make the bars slightly transparent by setting `alpha` to a small value."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "a8e9c378",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"drv\", fill=\"class\")) + geom_bar(alpha=0.5, position=\"identity\"))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "2aeffbf7",
+ "metadata": {},
+ "source": [
+ "The identity position adjustment is more useful for 2d geoms, like points, where it is the default.\n",
+ "\n",
+ "- `position = \"fill\"` works like stacking, but makes each set of stacked bars the same height.\n",
+ " This makes it easier to compare proportions across groups."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "14205000",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"drv\", fill=\"class\")) + geom_bar(position=\"fill\"))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "3a8e72bd",
+ "metadata": {},
+ "source": [
+ "- `position = \"dodge\"` places overlapping objects directly *beside* one another.\n",
+ " This makes it easier to compare individual values."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "c33c4a03",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"drv\", fill=\"class\")) + geom_bar(position=\"dodge\"))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "cd46f22b",
+ "metadata": {},
+ "source": [
+ "There's one other type of adjustment that's not useful for bar charts, but can be very useful for scatterplots.\n",
+ "Recall our first scatterplot.\n",
+ "Did you notice that the plot displays only some of the points (even though there are 234 observations in the dataset)?"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "ba4161de",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"displ\", y=\"hwy\")) + geom_point())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "28a3bfc7",
+ "metadata": {},
+ "source": [
+ "The underlying values of `hwy` and `displ` are rounded so the points appear on a grid and many points overlap each other.\n",
+ "This problem is known as **overplotting**.\n",
+ "This arrangement makes it difficult to see the distribution of the data.\n",
+ "Are the data points spread equally throughout the graph, or is there one special combination of `hwy` and `displ` that contains 109 values?\n",
+ "\n",
+ "You can avoid this gridding by setting the position adjustment to \"jitter\".\n",
+ "`position = \"jitter\"` adds a small amount of random noise to each point.\n",
+ "This spreads the points out because no two points are likely to receive the same amount of random noise."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "414ce7af",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"displ\", y=\"hwy\")) + geom_point(position=\"jitter\"))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "96277926",
+ "metadata": {},
+ "source": [
+ "Adding randomness seems like a strange way to improve your plot, but while it makes your graph less accurate at small scales, it makes your graph *more* revealing at large scales.\n",
+ "Because this is such a useful operation, ggplot2 comes with a shorthand for `geom_point(position = \"jitter\")`: `geom_jitter()`.\n",
+ "\n",
+ "Of course, a more sophisticated way of dealing with overplotting is via a binscatter plot, which is available in the [**binsreg**](https://nppackages.github.io/binsreg/) package.\n",
+ "\n",
+ "To learn more about position adjustment, take a look at the [documentation](https://lets-plot.org/)."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "90d09736",
+ "metadata": {},
+ "source": [
+ "### Exercises\n",
+ "\n",
+ "1. What is the problem with the following plot?\n",
+ " How could you improve it?"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "9bc38aef",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "(ggplot(mpg, aes(x=\"cty\", y=\"hwy\")) + geom_point())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "57b6523f",
+ "metadata": {},
+ "source": [
+ "2. What, if anything, is the difference between the two plots?\n",
+ " Why?\n",
+ "\n",
+ " ```python\n",
+ " (\n",
+ " ggplot(mpg, aes(x = \"displ\", y = \"hwy\")) +\n",
+ " geom_point()\n",
+ " )\n",
+ " (\n",
+ " ggplot(mpg, aes(x = \"displ\", y = \"hwy\")) +\n",
+ " geom_point(position = \"identity\")\n",
+ " )\n",
+ " ```"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "a3138f75",
+ "metadata": {},
+ "source": [
+ "3. What parameters to `geom_jitter()` control the amount of jittering?\n",
+ "\n",
+ "4. What's the default position adjustment for `geom_boxplot()`?\n",
+ " Create a visualisation of the `mpg` dataset that demonstrates it."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "7d302c31",
+ "metadata": {},
+ "source": [
+ "## The layered grammar of graphics\n",
+ "\n",
+ "We can expand on the graphing template you learned already by adding position adjustments, stats, coordinate systems, and faceting:\n",
+ "\n",
+ "```python\n",
+ "ggplot(data = ) + \n",
+ " (\n",
+ " mapping = aes(),\n",
+ " stat = , \n",
+ " position = \n",
+ " ) +\n",
+ " \n",
+ "```\n",
+ "\n",
+ "\n",
+ "Our new template takes six parameters, the bracketed words that appear in the template.\n",
+ "In practice, you rarely need to supply all seven parameters to make a graph because **lets-plot88 will provide useful defaults for everything except the data, the mappings, and the geom function.\n",
+ "\n",
+ "The six parameters in the template compose the grammar of graphics, a formal system for building plots.\n",
+ "The grammar of graphics is based on the insight that you can uniquely describe *any* plot as a combination of a dataset, a geom, a set of mappings, a stat, a position adjustment, a coordinate system, a faceting scheme, and a theme.\n",
+ "\n",
+ "To see how this works, consider how you could build a basic plot from scratch: you could start with a dataset and then transform it into the information that you want to display (with a stat).\n",
+ "Next, you could choose a geometric object to represent each observation in the transformed data.\n",
+ "You could then use the aesthetic properties of the geoms to represent variables in the data.\n",
+ "You would map the values of each variable to the levels of an aesthetic.\n",
+ "These steps are illustrated in the figure below.\n",
+ "\n",
+ "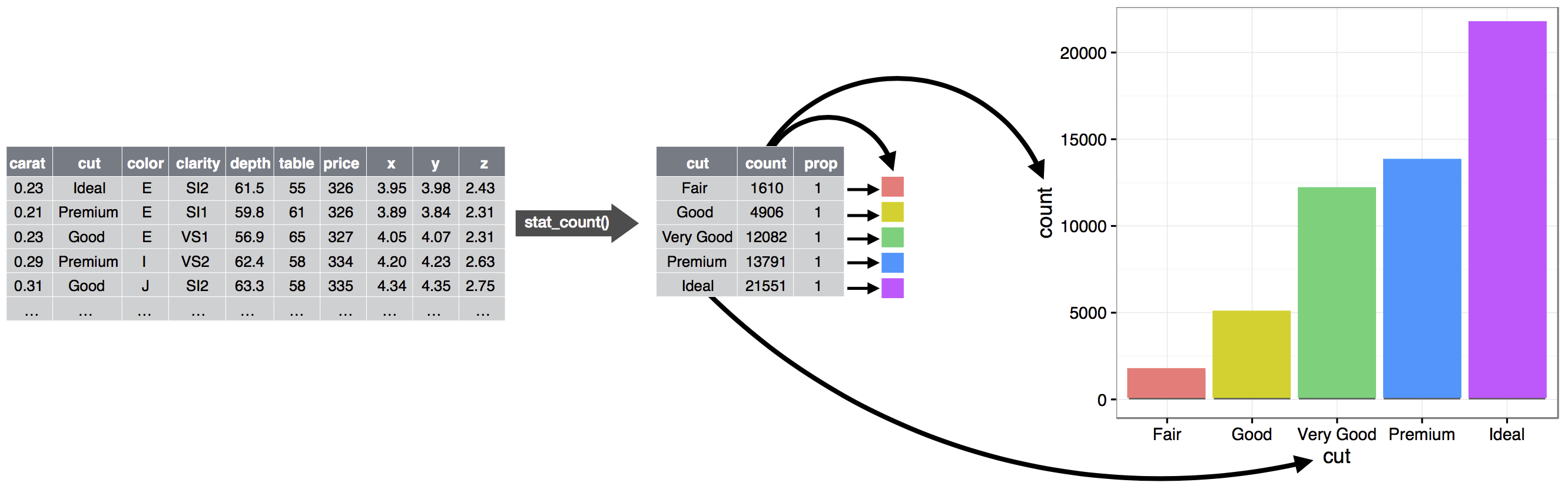"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "c0f7a7ff",
+ "metadata": {},
+ "source": [
+ "At this point, you would have a complete graph, but you could further adjust the positions of the geoms within the coordinate system (a position adjustment) or split the graph into subplots (faceting).\n",
+ "You could also extend the plot by adding one or more additional layers, where each additional layer uses a dataset, a geom, a set of mappings, a stat, and a position adjustment.\n",
+ "\n",
+ "You could use this method to create a lot of plots that you can imagine.\n",
+ "\n",
+ "## Summary\n",
+ "\n",
+ "In this chapter you learned about the layered grammar of graphics starting with aesthetics and geometries to build a simple plot, facets for splitting the plot into subsets, statistics for understanding how geoms are calculated, position adjustments for controlling the fine details of position when geoms might otherwise overlap, and coordinate systems which allow you to fundamentally change what `x` and `y` mean.\n",
+ "\n",
+ "The most useful further resource on **lets-plot** is the documentation, which you [can find here](https://lets-plot.org/)."
+ ]
+ }
+ ],
+ "metadata": {
+ "interpreter": {
+ "hash": "9d7534ecd9fbc7d385378f8400cf4d6cb9c6175408a574f1c99c5269f08771cc"
+ },
+ "jupytext": {
+ "cell_metadata_filter": "-all",
+ "encoding": "# -*- coding: utf-8 -*-",
+ "formats": "md:myst",
+ "main_language": "python"
+ },
+ "kernelspec": {
+ "display_name": "Python 3 (ipykernel)",
+ "language": "python",
+ "name": "python3"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.10.12"
+ },
+ "toc-showtags": true
+ },
+ "nbformat": 4,
+ "nbformat_minor": 5
+}
diff --git a/visualise.md b/visualise.md
new file mode 100644
index 0000000..3e2f59f
--- /dev/null
+++ b/visualise.md
@@ -0,0 +1,120 @@
+---
+jupytext:
+ cell_metadata_filter: -all
+ formats: md:myst
+ text_representation:
+ extension: .md
+ format_name: myst
+ format_version: '0.8'
+ jupytext_version: 1.5.0
+kernelspec:
+ display_name: 'Python 3.10.12 64-bit (''py4ds2e'': conda)'
+ language: python
+ name: python3
+---
+(visualise)=
+# Visualisation
+
+After reading the first part of the book, you understand the basics of the most important tools for doing data science. Now it’s time to start diving into the details. In this part of the book, you’ll learn about visualising data in further depth (in {ref}`vis-layers`), and get further stuck into the details of the different kinds of data visualisation (in {ref}`exploratory-data-analysis` and {ref}`communicate-plots`). In this short chapter, we discuss the different ways to create visualisations, and the different purposes of visualisations.
+
+## Philosophies of data visualisation
+
+There are broadly two categories of approach to using code to create data visualisations: *imperative* (build what you want from individual elements) and *declarative* (say what you want from a list of pre-existing options). Choosing which to use involves a trade-off: imperative libraries offer you flexibility but at the cost of some verbosity; declarative libraries offer you a quick way to plot your data, but only if it’s in the right format to begin with, and customisation to special chart types is more difficult.
+
+Python has many excellent plotting packages, including perhaps the most powerful imperative plotting package around, **matplotlib**, and an amazing declarative library that we already saw, **lets-plot**. These two libraries will get you a long way, and each could be worthy of an entire book themselves. Fortunately for us, though, we can do 95% of what we need with a small number of commands from one or the other of them. In general, to keep this book as light as possible, we've opted to use **lets-plot** wherever possible—and {ref}`vis-layers` is going to take you on a more in-depth tour of how to use it yourself.
+
+## Purposes of data visualisation
+
+Data visualisation has all kinds of different purposes. It can be useful to bear in mind three broad categories of visualisation that are out there:
+
+- exploratory
+- scientific
+- narrative
+
+Let's look at each in a bit more detail.
+
+### Exploratory Data Viz
+
+The first of the three kinds is *exploratory data visualisation*, and it's the kind that you do when you're looking and data and trying to understand it. Just plotting the data is a really good strategy for getting a feel for any issues there might be. This is perhaps most famously demonstrated by Anscombe's quartet: four different datasets with the same mean, standard deviation, and correlation but very different data distributions.
+
+```{code-cell} ipython3
+---
+tags: [remove-input]
+---
+import numpy as np
+import pandas as pd
+import matplotlib.pyplot as plt
+import matplotlib_inline.backend_inline
+
+# Plot settings
+plt.style.use("https://github.com/aeturrell/python4DS/raw/main/plot_style.txt")
+matplotlib_inline.backend_inline.set_matplotlib_formats("svg")
+
+# Set max rows displayed for readability
+pd.set_option("display.max_rows", 6)
+
+x = [10, 8, 13, 9, 11, 14, 6, 4, 12, 7, 5]
+y1 = [8.04, 6.95, 7.58, 8.81, 8.33, 9.96, 7.24, 4.26, 10.84, 4.82, 5.68]
+y2 = [9.14, 8.14, 8.74, 8.77, 9.26, 8.10, 6.13, 3.10, 9.13, 7.26, 4.74]
+y3 = [7.46, 6.77, 12.74, 7.11, 7.81, 8.84, 6.08, 5.39, 8.15, 6.42, 5.73]
+x4 = [8, 8, 8, 8, 8, 8, 8, 19, 8, 8, 8]
+y4 = [6.58, 5.76, 7.71, 8.84, 8.47, 7.04, 5.25, 12.50, 5.56, 7.91, 6.89]
+
+datasets = {"I": (x, y1), "II": (x, y2), "III": (x, y3), "IV": (x4, y4)}
+
+fig, axs = plt.subplots(
+ 2,
+ 2,
+ sharex=True,
+ sharey=True,
+ figsize=(10, 6),
+ gridspec_kw={"wspace": 0.08, "hspace": 0.08},
+)
+axs[0, 0].set(xlim=(0, 20), ylim=(2, 14))
+axs[0, 0].set(xticks=(0, 10, 20), yticks=(4, 8, 12))
+
+for ax, (label, (x, y)) in zip(axs.flat, datasets.items()):
+ ax.text(0.1, 0.9, label, fontsize=20, transform=ax.transAxes, va="top")
+ ax.tick_params(direction="in", top=True, right=True)
+ ax.plot(x, y, "o")
+
+ # linear regression
+ p1, p0 = np.polyfit(x, y, deg=1) # slope, intercept
+ ax.axline(xy1=(0, p0), slope=p1, color="r", lw=2)
+
+ # add text box for the statistics
+ stats = (
+ f"$\\mu$ = {np.mean(y):.2f}\n"
+ f"$\\sigma$ = {np.std(y):.2f}\n"
+ f"$r$ = {np.corrcoef(x, y)[0][1]:.2f}"
+ )
+ bbox = dict(boxstyle="round", fc="blanchedalmond", ec="orange", alpha=0.5)
+ ax.text(
+ 0.95,
+ 0.07,
+ stats,
+ fontsize=9,
+ bbox=bbox,
+ transform=ax.transAxes,
+ horizontalalignment="right",
+ )
+
+plt.suptitle("Anscombe's Quartet")
+plt.show()
+```
+
+Exploratory visualisation is usually quick and dirty, and flexible too. Some exploratory data viz can be automated, and there's a whole host of packages to help with this, including [**skimpy**](https://aeturrell.github.io/skimpy/).
+
+Beyond you and perhaps your co-authors/collaborators, however, not many other people should be seeing your exploratory visualisation! They will typically be worked up quickly, be numerous, and be throw-away. We'll look more at this in {ref}`exploratory-data-analysis`.
+
+### Scientific Data Viz
+
+The second kind, scientific data visualisation, is the prime cut of your exploratory visualisation. It's the kind of plot you might include in a more technical paper, the picture that says a thousand words. I often think of the first image of a black hole {cite:t}`akiyama2019first` as a prime example of this. You can get away with having a high density of information in a scientific plot and, in short format journals, you may need to. The journal Physical Review Letters, which has an 8 page limit, has a classic of this genre in more or less every issue. Ensuring that important values can be accurately read from the plot is especially important in these kinds of charts. But they can also be the kind of plot that presents the killer results in a study; they might not be exciting to people who don't look at charts for a living, but they might be exciting and, just as importantly, understandable by your peers.
+
+This type of visualisation is especially popular in the big science journals like *Nature* and *Science*, where space is at a premium. We won't cover this type of plot in this book, because it tends to be very bespoke.
+
+### Narrative Data Viz
+
+The third and final kind is narrative data visualisation. This is the one that requires the most thought in the step where you go from the first view to the end product. It's a visualisation that doesn't just show a picture, but gives an insight. These are the kind of visualisations that you might see in the *Financial Times*, *The Economist*, or on the *BBC News* website. They come with aids that help the viewer focus on the aspects that the creator wanted them to (you can think of these aids or focuses as doing for visualisation what bold font does for text). They're well worth using in your work, especially if you're trying to communicate a particular narrative, and especially if the people you're communicating with don't have deep knowledge of the topic. You might use them in a paper that you hope will have a wide readership, in a blog post summarising your work, or in a report intended for a policymaker.
+
+You can find more information on the topic of communicating via data visualisations in the {ref}`communicate-plots` chapter.
diff --git a/workflow-basics.ipynb b/workflow-basics.ipynb
index 9d17886..c7ac6b5 100644
--- a/workflow-basics.ipynb
+++ b/workflow-basics.ipynb
@@ -42,7 +42,7 @@
"id": "0f0ee026",
"metadata": {},
"source": [
- "The extra package **numpy** contains many of the additional mathematical operators that you might need. If you don't already have **numpy** installed, open up the terminal in Visual Studio Code (go to \"Terminal -> New Terminal\" and then type `conda install numpy` into the terminal then hit return). Once you have **numpy** installed, you can import it and use it like this:"
+ "The extra package **numpy** contains many of the additional mathematical operators that you might need. If you don't already have **numpy** installed, open up the terminal in Visual Studio Code (go to \"Terminal -> New Terminal\" and then type `pip install numpy` into the terminal then hit return). Once you have **numpy** installed, you can import it and use it like this:"
]
},
{
@@ -172,7 +172,13 @@
"id": "cb49df46",
"metadata": {},
"source": [
- "With short pieces of code like this, it is not necessary to leave a command for every single line of code and you should try to use informative names wherever you can because these help readers of your code (likely to be you in the future) understand what is going on!"
+ "With short pieces of code like this, it is not necessary to leave a command for every single line of code and you should try to use informative names wherever you can because these help readers of your code (likely to be you in the future) understand what is going on!\n",
+ "\n",
+ "Our advice is to use comments to explain the *why* of your code, not the *how* or the *what*. The *what* and *how* of your code are always possible to figure out, even if it might be tedious, by carefully reading it. If you describe every step in the comments, and then change the code, you will have to remember to update the comments as well (tedious) or it will be confusing when you return to your code in the future.\n",
+ "\n",
+ "Figuring out *why* something was done is much more difficult, if not impossible. For example, geom_smooth() has an argument called span, that controls the smoothness of the curve, with larger values yielding a smoother curve. Suppose you decide to change the value of span from its default of 0.75 to 0.9: it’s easy for a future reader to understand *what* is happening, but unless you note your thinking in a comment, no one will understand *why* you changed the default.\n",
+ "\n",
+ "For data analysis code, use comments to explain your overall plan of attack and record important insights as you encounter them. There’s no way to re-capture this knowledge from the code itself."
]
},
{
@@ -244,7 +250,7 @@
"\n",
"Object names in Python are case-sensitive too, so `age`, `Age` and `AGE` could all be three different variables.\n",
"\n",
- "When you're naming objects, it's best to make them descriptive so you can keep track of what they are. You’ll need to adopt a convention for multiple words. We recommend snake_case, where you separate lowercase words with `_`. For example, `i_use_snake_case` could be an object name.\n"
+ "When you're naming objects, it's best to make them descriptive so you can keep track of what they are. You’ll need to adopt a convention for multiple words. We recommend snake_case, where you separate lowercase words with `_`. For example, `i_use_snake_case` is a valid snake case name for an object.\n"
]
},
{
@@ -258,6 +264,97 @@
"```"
]
},
+ {
+ "cell_type": "markdown",
+ "id": "4d364f79",
+ "metadata": {},
+ "source": [
+ "Remember that you can always inspect an object that you've created by typing its name again:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "a5998cb5",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "primes"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "82807f98",
+ "metadata": {},
+ "source": [
+ "Make another assignment:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "ae55d78e",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "this_is_a_really_long_name = 2.5"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "73c37168",
+ "metadata": {},
+ "source": [
+ "To save yourself time in inspecting this object via the interactive window, you can just begin typing the name (type \"this\") and then hit the TAB button. Visual Studio Code will autocomplete what you've written using the variables you've defined during your session. This is a top tip to save time!\n",
+ "\n",
+ "If you're using the interactive console, rather than a notebook, there's another top tip. Let's say you previously ran `this_is_a_really_long_name = 2.5` but you *meant* to set it to 3.5. Don't despair; you don't have to type it all out again. With your cursor in the interactive window, you can simply hit ↑ on your keyboard and cycle through previous commands you issued. Change 2.5 to 3.5, hit shift + return, and you'll have redefined your variable.\n",
+ "\n",
+ "Let's define another variable:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "038c7d52",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "py_variable = 2 ^ 3"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "446dfa44",
+ "metadata": {},
+ "source": [
+ "Now let's try to inspect it:"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "5248dbc8",
+ "metadata": {},
+ "source": [
+ "```python\n",
+ "py_variabl\n",
+ "\n",
+ "---------------------------------------------------------------------------\n",
+ "NameError Traceback (most recent call last)\n",
+ "/Users/aet/Documents/git_projects/python4DS/workflow-basics.ipynb Cell 31 in ()\n",
+ "----> 1 py_variabl\n",
+ "\n",
+ "NameError: name 'py_variabl' is not defined\n",
+ "```"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "e2cf9efc",
+ "metadata": {},
+ "source": [
+ "This illustrates the brilliance and frustration of coding: your IDE (Visual Studio Code) will do tedious computations for you, but, in exchange, you must be precise in your instructions. If not, you’re likely to get an error that says the object you’re looking for was not found. Typos matter; Python can’t read your mind and say, “oh, they probably meant `py_variable` when they typed `py_variabl`”."
+ ]
+ },
{
"attachments": {},
"cell_type": "markdown",
@@ -330,22 +427,80 @@
]
},
{
- "attachments": {},
"cell_type": "markdown",
- "id": "51346af1",
+ "id": "d380a16b",
"metadata": {},
"source": [
- "````{admonition} Exercise\n",
- "\n",
- "Why does this code not work?\n",
+ "Just as with variables, code completion works on functions too. Try typing in `su` and hitting tab to see this in action."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "0e578077",
+ "metadata": {},
+ "source": [
+ "You'll need to be extra careful with objects that are strings (words, sentences, letters, and phrases), because these always need to come with quotation marks around them. You can use single or double quotation marks as you like, but i) the convention is double quotation marks, and ii) it's good to be consistent, whichever you choose. \n",
"\n",
+ "Here's an example of some code that throws an error"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "1b95d213",
+ "metadata": {},
+ "source": [
"```python\n",
- "my_variable = 10\n",
- "my_varıable\n",
- "```\n",
+ "x = \"hello\n",
+ "\n",
+ " Input In [3]\n",
+ " x = \"hello\n",
+ " ^\n",
+ "SyntaxError: unterminated string literal (detected at line 1)\n",
+ "```"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "38d073ce",
+ "metadata": {},
+ "source": [
+ "Again, Visual Studio Code can really help you out here because as soon as you open a double quotation mark, it will have the closing one ready for you."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "id": "6af6ea6c",
+ "metadata": {},
+ "source": [
+ "## Exercises\n",
+ "\n",
+ "1. Why does this code not work?\n",
+ "\n",
+ " ```python\n",
+ " my_variable = 10\n",
+ " my_varıable\n",
+ " ```\n",
+ "\n",
+ " Look carefully! This may seem like an exercise in pointlessness, but training your brain to notice even the tiniest difference will pay off when programming.\n",
+ "\n",
+ "2. Tweak each of the following Python commands so that they run correctly: \n",
+ "\n",
+ " ```python\n",
+ " import pandas as pd\n",
+ " from palmerpenguins import load_penguins\n",
+ " from lets_pot import *\n",
+ "\n",
+ " LetsPlot.setup_html()\n",
+ " penguins = load_penguins()\n",
"\n",
- "Look carefully! This may seem like an exercise in pointlessness, but training your brain to notice even the tiniest difference will pay off when programming.\n",
- "````"
+ " (\n",
+ " ggplot(\n",
+ " dTA=penguins,\n",
+ " maping=aes(x=\"flipper_length_mm\", y=\"body_mass_g\", color=\"species\"),\n",
+ " )\n",
+ " + geom_smooth(method=\"lm)\n",
+ " )\n",
+ " ```"
]
}
],
@@ -374,7 +529,7 @@
"name": "python",
"nbconvert_exporter": "python",
"pygments_lexer": "ipython3",
- "version": "3.9.12"
+ "version": "3.10.12"
},
"toc-showtags": true
},
diff --git a/workflow-help.md b/workflow-help.md
index 45ae4cb..48f05d5 100644
--- a/workflow-help.md
+++ b/workflow-help.md
@@ -14,15 +14,29 @@ kernelspec:
This book is not an island; there is no single resource that will allow you to master Python for Data Science. As you begin to apply the techniques described in this book to your own data, you will soon find questions that we do not answer. This section describes a few tips on how to get help, and to help you keep learning.
+## Resources
+
+Some other resources for learning are:
+
+- [The Python Data Science Handbook](https://jakevdp.github.io/PythonDataScienceHandbook/)
+- [Real Python](https://realpython.com/), which has excellent short tutorials that cover Python more broadly (not just data science)
+- [freeCodeCamp's Python courses](https://www.freecodecamp.org/news/search?query=data%20science%20python), though take care to select one that's at the right level for you
+- [Coding for Economists](https://aeturrell.github.io/coding-for-economists), which has similar content to this book but is more in depth and aimed at analysts (particularly in economics)
+
## Google is your friend
If you get stuck, start with Google. Typically adding "Python" or "Python Data Science" (as the Python ecosystem goes *well* beyond data science) to a query is enough to restrict it to relevant results. Google is particularly useful for error messages. If you get an error message and you have no idea what it means, try googling it! Chances are that someone else has been confused by it in the past, and there will be help somewhere on the web.
If Google doesn't help, try [Stack Overflow](http://stackoverflow.com). Start by spending a little time searching for an existing answer, including `[Python]` to restrict your search to questions and answers that use Python.
+## In the loop
+
+It's also helpful to keep an eye on the latest developments in data science. There are tons of data science newsletters out there, and we recommend keeping up with the Python data science community by following the (#pydata), (#datascience), and (#python) hashtags on Twitter.
+
## Making a reprex (reproducible example)
If your googling doesn't find anything useful, it's a really good idea prepare a minimal reproducible example or **reprex**.
+
A good reprex makes it easier for other people to help you, and often you'll figure out the problem yourself in the course of making it. There are two parts to creating a reprex:
- First, you need to make your code reproducible. This means that you need to capture everything, i.e., include any packages you used and create all necessary objects. The easiest way to make sure you've done this is to use the [**watermark**](https://github.com/rasbt/watermark) package alongside whatever else you are doing:
@@ -66,7 +80,3 @@ df
- **Code**: copy and paste the minimal reproducible example code (including the packages, as noted above). Make sure you've used spaces and your variable names are concise, yet informative. Use comments to indicate where your problem lies. Do your best to remove everything that is not related to the problem. Finally, the shorter your code is, the easier it is to understand, and the easier it is to fix.
Finish by checking that you have actually made a reproducible example by starting a fresh Python session and copying and pasting your reprex in.
-
-## Investing in yourself
-
-You should also spend some time preparing yourself to solve problems before they occur. Investing a little time in learning Python each day will pay off in the long run!