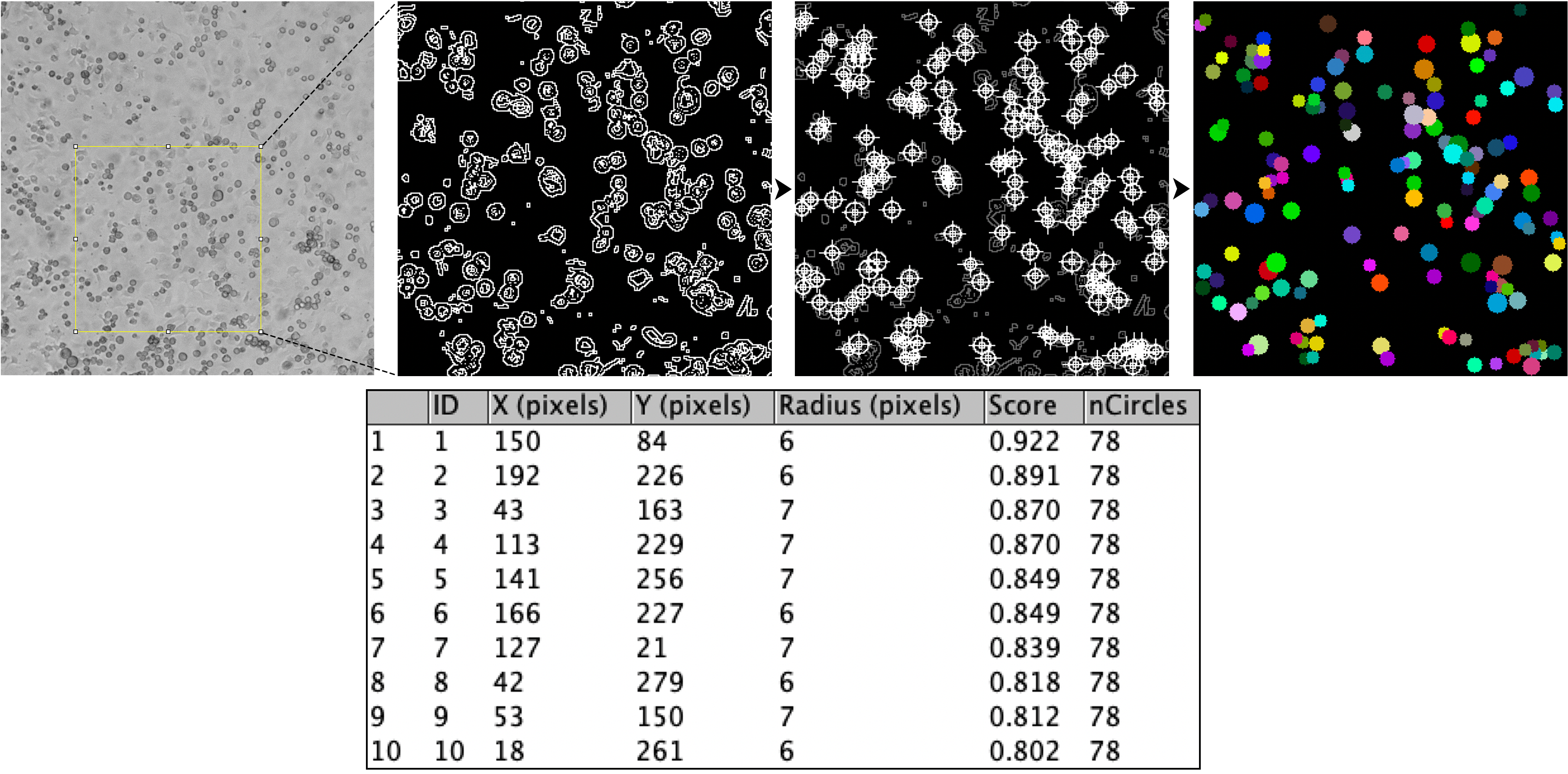
Automated quantification of cell number, radius, and area in culture using randomized ROI selection and hugh circles.
Must have Hough Circle Transform Installed
link: https://imagej.net/plugins/hough-circle-transform
Manual:
Open brightfield image in Fiji
Code:
Insert code into File --> New --> Script...
Change Language --> Python
Random ROI selection --> Find Edges --> Auto Threshold --> Mask Conversion --> Binary Outline --> Hugh Circle Transform --> Data in Results Table
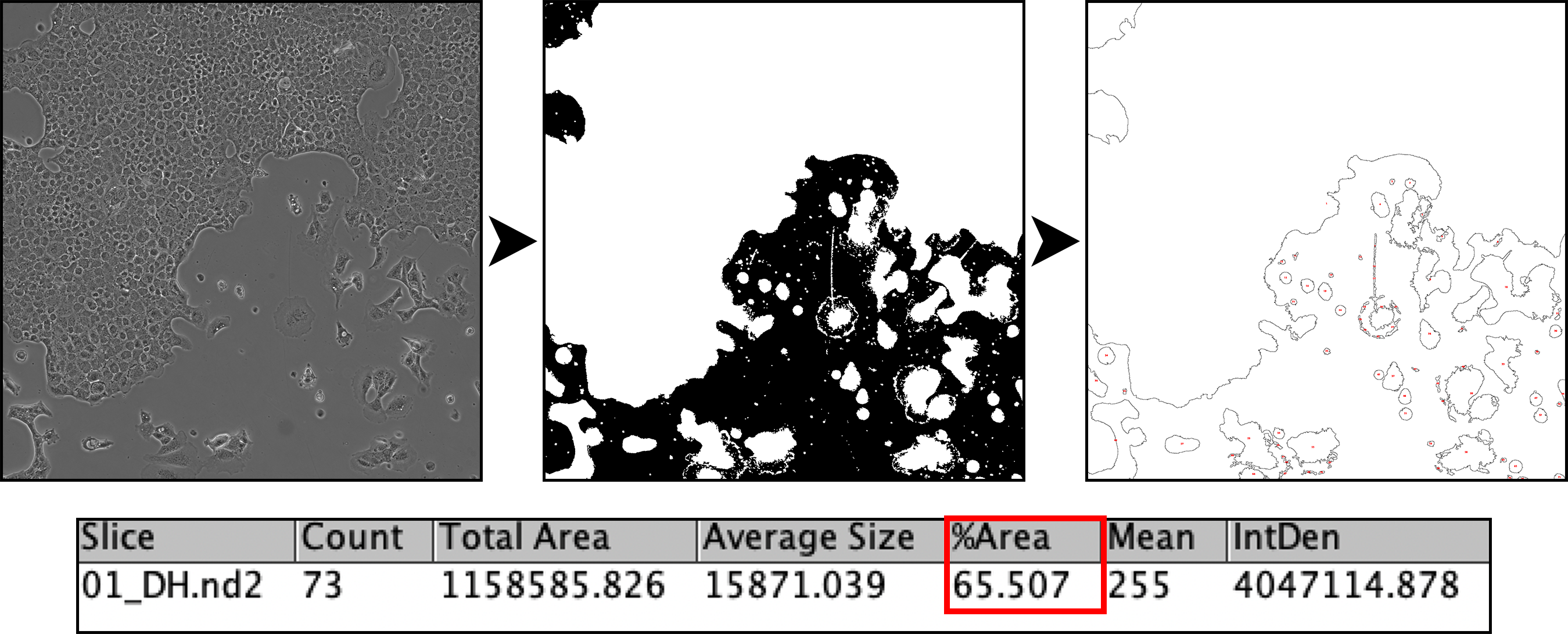
Automated quantification of confluency (%area) in culture using binary masking, and particle analysis.
Manual:
Open brightfield image in Fiji
Code:
Insert code into File --> New --> Script...
Change Language --> Python
Find Edges --> Custom (set) Threshold --> Mask Conversion --> Dilate --> Fill Holes --> Analyze particles (custom)--> Data in Results Table
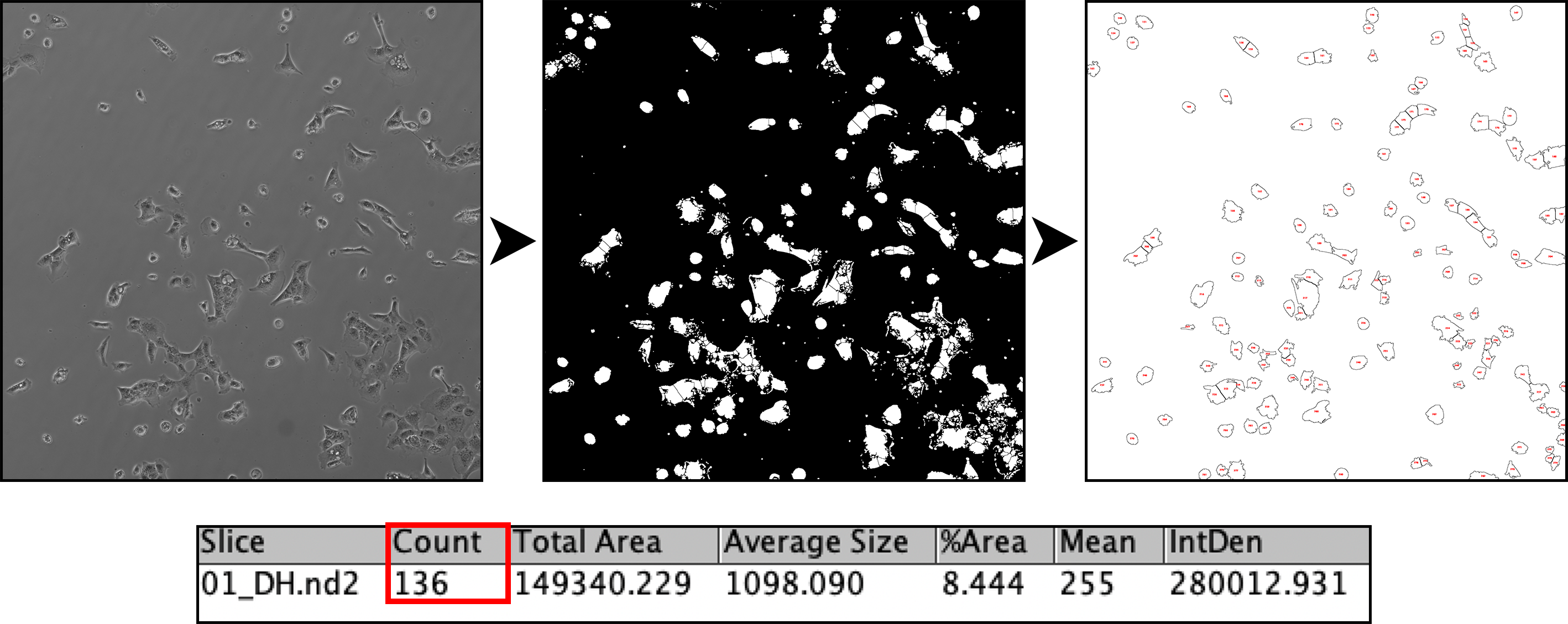
Automated quantification of cell number in culture using binary masking, and particle analysis.
Manual:
Open brightfield image in Fiji
Code:
Insert code into File --> New --> Script...
Change Language --> Python
Find Edges --> Mask Conversion --> Dilate --> Fill Holes --> Watershed --> Analyze particles (custom) --> Data in Results Table