This repository contains two things:
- slides.pdf (Slides from the talk)
- example-project (showcasing the various tools that I described in the talk)
To try things out in the example project, clone this repository (click the green button above that says Code). Then try out devtools/git/Snakemake.
Open MyProject.Rproj in RStudio (RStudio isn't necessary for this, but I assume that's what most people use). Then run:
devtools::load_all()or use the keyboard shortcut Ctrl+Shift+L. Type ?month_merge and the documentation should show up. Then run the example (which you'll be able to do since devtools has loaded all the project-specific functions).
You can run all the examples with
devtools::run_examples()If you want, play around with editing any of the functions, re-loading, and seeing that everything stays correctly sourced. Note that if you change the documentation, you'll need to rebuild it with devtools::document() or Ctrl+Shift+D.
As I say in the slides, I can't cover all of what git can do here. But to show you how it can help reduce clutter, just click on example-project -> R -> month-merge.r -> History, then "View code at this point" for some past commit to see how everything looked in an earlier version.
Or try deleting month-merge.r on your machine, then running
git checkout R/month-merge.rHey! It's back! This should give you much more peace of mind about deleting things.
Note: Snakemake is a Python package, so you have to install it using pip or whatever you use to install Python packages.
Once you've installed Snakemake, use it to create an output target. Here's a table of regression results (note: depending on your setup, you may need to submit snakemake --cores 1 output/results.html):
snakemake output/results.htmlNote that Snakemake knows that the regression results depend on running multiple imputation first, so it does both. If you run
snakemake output/results.htmlagain, note that nothing happens because Snakemake knows the results are up to date. Now try deleting output/results.html and running
snakemake output/results.htmlNote that Snakemake creates the table, but doesn't re-run the imputation step because it doesn't need to. You can play around with deleting various files in output and see what happens, or changing the code in the scripts, and see how Snakemake recognizes what steps need to be re-run.
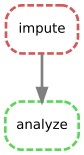
Lastly, a neat thing is that Snakemake can automatically make a DAG for you that shows you how your rules are related to each other:
snakemake output/results.html --dag | dot | displaywhich in this case produces
It's not super exciting for this simple project but you can imagine how helpful this would be in more complex scenarios.
More information on the packages mentioned in the slides: