-
Notifications
You must be signed in to change notification settings - Fork 823
Closed
Description
Expected behavior
FastNS misses neighbors in a perfect 111 FCC slab (an ideal, oversimplified case).
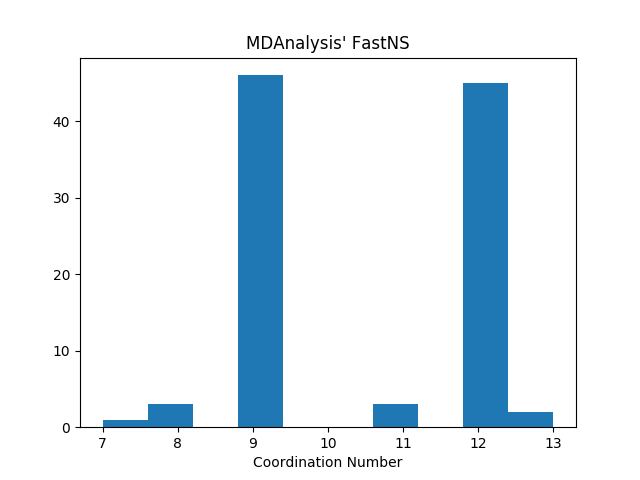
Actual behavior
Instead of identifying 9 or 12 neighbors for every atom, FastNS identifies 9 or 12 for most atoms, with some strange exceptions:
Or see the structure, with atoms colored by coordination number:
Code to reproduce the behavior
(Requires ase.)
The Ag-Ag distance in this system is 2.89Å, so a 3.2Å cutoff is quite sufficient.
from ase.build import fcc111
from MDAnalysis.lib.mdamath import triclinic_box
from MDAnalysis.lib.nsgrid import FastNS
slab = fcc111('Ag', size = (5, 5, 4), vacuum = 10.0)
tcbox = triclinic_box(*slab.cell)
fns = FastNS(cutoff = 3.2, coords = slab.positions, box = tcbox, pbc = True)
ns = fns.self_search()
cns = [len(n) for n in ns.get_indices()]
print(np.unique(cns, return_counts = True))
# => (array([ 7, 8, 9, 11, 12, 13]), array([ 1, 3, 46, 3, 45, 2]))The results are identical for cutoffs of 3.0Å and 2.9Å. At a cutoff of 2.5Å, as expected, all atoms have a CN of 0.
The results are correct, for example, using pymatgen's (very, very, very slow) CutOffDictNN:
from pymatgen.io.ase import AseAtomsAdaptor
from pymatgen.analysis.local_env import CutOffDictNN
cdnn = CutOffDictNN(cut_off_dict={('Ag', 'Ag') : 3.0})
slabstruct = AseAtomsAdaptor.get_structure(slab)
cs_matcutoff = [len(e) for e in cdnn.get_all_nn_info(slabstruct)]
print(np.unique(cs_matcutoff, return_counts = True))
# => (array([ 9, 12]), array([50, 50]))Currently version of MDAnalysis
- Which version are you using? 0.19.2
- Which version of Python (
python -V)? Python 3.6.8 :: Anaconda, Inc. - Which operating system? Linux (Ubuntu)
Reactions are currently unavailable