Added 2 algorithms to solve LASSO-like problems: restarting FISTA and POGM'#7
Conversation
|
I guess since the tests are passing, the conda install method is fine. |
|
any plot you can share?
|
|
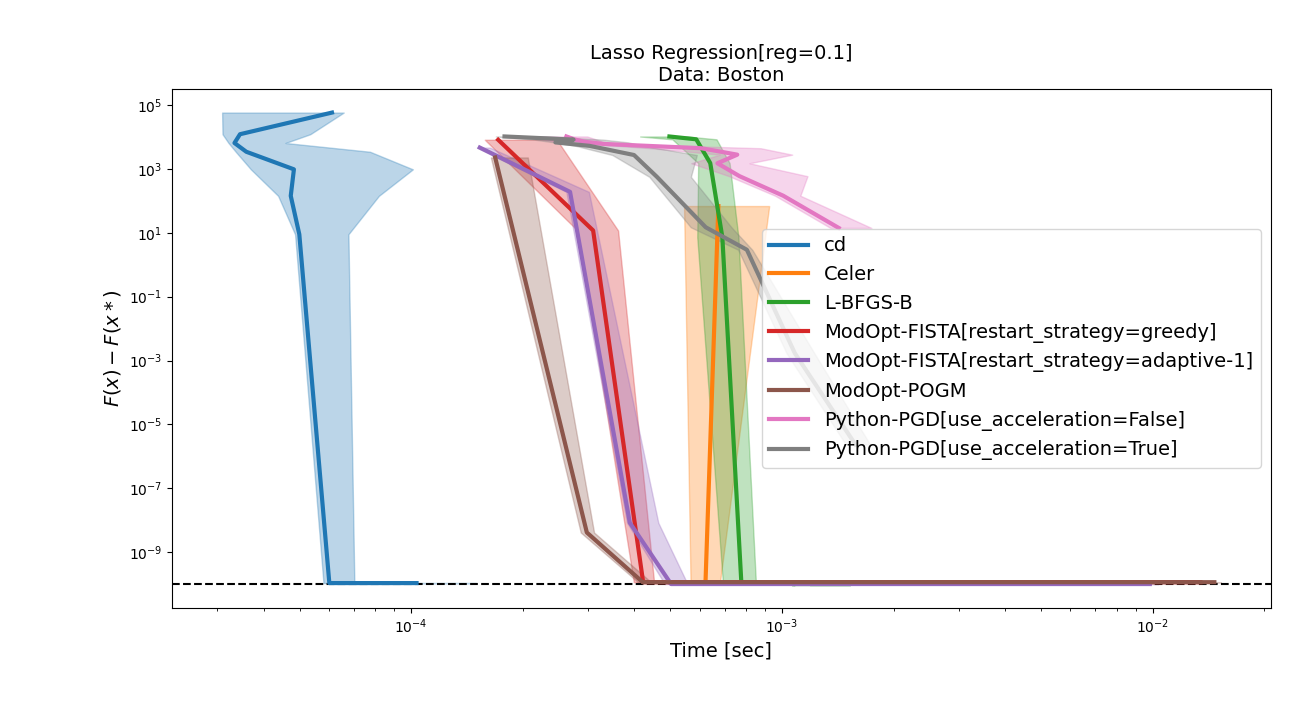
Sure which ones do you have in mind? Here is for example the results on the Boston dataset against CD, Celer, L-BFGS-B and PGD, with The simulated ones are quite slow (and fista greedy diverges with the current hyperparameter setup in some cases), but I can run them if they make things clearer. |
|
yes it's slow on simulated.
what you mean it diverges?
… |
|
Well FISTA greedy uses a strategy where the step size begins by being superior to the inverse of the Lipschitz constant. There is a parameter telling you by how much it should be bigger (in this case 1.3x). Here, I didn't want to tune the hyperparameters, so I set it at the recommended value which is supposed to work well in most settings (like for MRI reconstruction, on the Boston dataset, or some simulated cases). Unfortunately for some cases of simulated data, it doesn't and the algorithm diverges. Re simulated: do you want me to run the benchmarks or is the boston one enough? |
|
ok got it
I am not requesting anything. I am just curious :)
… |
 tomMoral
left a comment
tomMoral
left a comment
There was a problem hiding this comment.
A small comment on the initialization of the method. To be consistent with other, you need to compute L in the run.
|
@tomMoral Done ! (The reason it doesn't seem very natural code-wise is because those 2 algorithms were implemented in the spirit of the inverse problem framework, where the If I may, I noticed that one another algorithm made a computation before |
 tomMoral
left a comment
tomMoral
left a comment
There was a problem hiding this comment.
Sorry for the long delay @zaccharieramzi !!
LGTM! merging
|
For the matrix vector product in glmnet, this is a bit different because this our way to set the lambda parameter for this solver to the same value as for the others, not something that is necessary for the solver. So I would say this is correct as is but I am ready to discuss if you think otherwise. |
These 2 algorithms are implemented in ModOpt, where they were intended to solve inverse problems (this is why the notations might seem a bit funny).
The defaults are those from this paper (that I wrote at the beginning of my thesis so please don't be mean).
I saw that none of the other solvers had docs about what they were implementing (i.e. a citation) or their potential hyperparameters, so I didn't do it for those 2, but happy to do it. Otherwise, all there is to know is inside the ModOpt package.
I still need to check that the conda install method is working.
Should I post some results on boston and simulated datasets here?