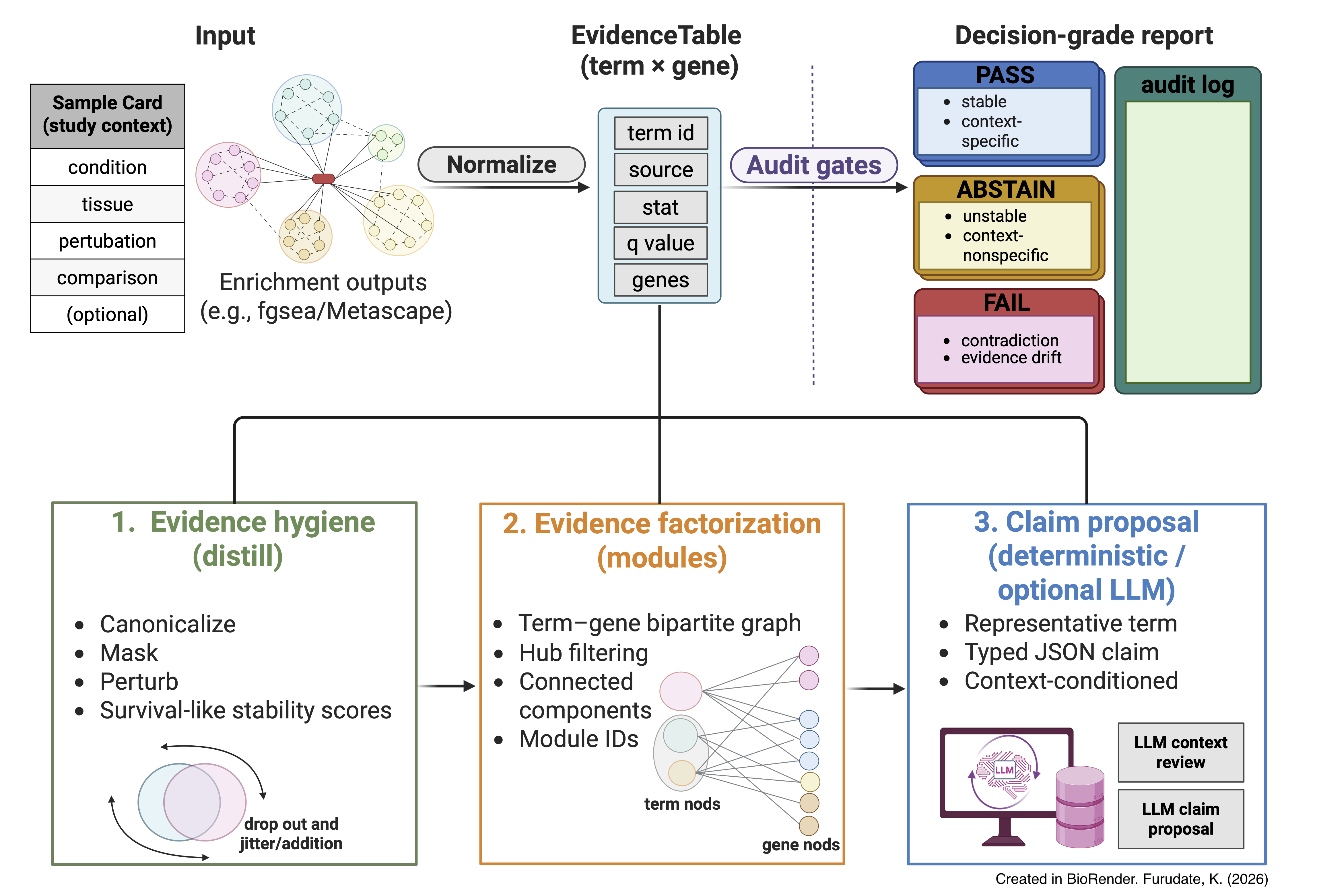
Enrichment interpretations → audit-gated, decision-grade claims.
Enrichment interpretations → audit-gated, decision-grade claims.
- Docs: https://llm-pathwaycurator.readthedocs.io/
- Paper reproducibility (canonical):
paper/(seepaper/README.md; panel map inpaper/FIGURE_MAP.csv) - Cite: bioRxiv preprint (DOI: 10.64898/2026.02.18.706381).
LLM-PathwayCurator is an interpretation quality-assurance (QA) layer for enrichment analysis.
It does not introduce a new enrichment statistic. Instead, it turns EA outputs into auditable decision objects:
- Input: enrichment term lists from ORA (e.g., Metascape) or rank-based enrichment (e.g., fgsea, an implementation of the GSEA method)
- Output: typed, evidence-linked claims + PASS/ABSTAIN/FAIL decisions + reason-coded audit logs
- Promise: we abstain when claims are unstable, under-supported, contradictory, or context-nonspecific
Selective prediction for pathway interpretation: calibrated abstention is a feature, not a failure.
Fig. 1a. Overview of LLM-PathwayCurator workflow:
EvidenceTable → modules → claims → audits
(bioRxiv preprint)
Enrichment tools return ranked term lists. In practice, interpretation breaks because:
- Representative terms are ambiguous under study context
- Gene support is opaque, enabling cherry-picking
- Related terms share / bridge evidence in non-obvious ways
- There is no mechanical stop condition for fragile narratives
LLM-PathwayCurator replaces narrative endorsement with audit-gated decisions.
We transform ranked terms into machine-auditable claims by enforcing:
- Evidence-linked constraints: claims must resolve to valid term/module identifiers and supporting-gene evidence
- Stability audits: supporting-gene perturbations yield stability proxies (operating point: τ)
- Context validity stress tests: context swap reveals context dependence without external knowledge
- Contradiction checks: internally inconsistent claims fail mechanically
- Reason-coded outcomes: every decision is explainable by a finite audit code set
- Not an enrichment method; it audits enrichment outputs.
- Not a free-text summarizer; claims are schema-bounded (typed JSON; no narrative prose as “evidence”).
- Not a biological truth oracle; it checks internal consistency and evidence integrity, not mechanistic truth.
A) Stability distillation (evidence hygiene)
Perturb supporting genes (seeded) to compute stability proxies (e.g., LOO/jackknife-like survival scores).
Output: distilled.tsv
B) Evidence factorization (modules)
Factorize the term–gene bipartite graph into evidence modules that preserve shared vs distinct support.
Outputs: modules.tsv, term_modules.tsv, term_gene_edges.tsv
C) Claims → audit → report
- C1 (proposal-only): deterministic baseline or optional LLM proposes typed claims with resolvable evidence links
- C2 (audit/decider): mechanical rules assign PASS/ABSTAIN/FAIL with precedence (FAIL > ABSTAIN > PASS)
- C3 (report): decision-grade report + audit log (
audit_log.tsv) + provenance
llm-pathway-curator run \
--sample-card examples/demo/sample_card.json \
--evidence-table examples/demo/evidence_table.tsv \
--out out/demo/audit_log.tsv— PASS/ABSTAIN/FAIL + reason codes (mechanical)report.jsonl,report.md— decision objects (evidence-linked)claims.proposed.tsv— proposed candidates (proposal-only; auditable)distilled.tsv— stability proxies / evidence hygiene outputsmodules.tsv,term_modules.tsv,term_gene_edges.tsv— evidence structurerun_meta.json(+ optionalmanifest.json) — pinned params + provenance
LLM-PathwayCurator includes two small post-processing commands for ranking and publication-ready visualization of ranked terms/modules:
llm-pathway-curator rank— produces a ranked table (claims_ranked.tsv) for downstream plots and summaries.llm-pathway-curator plot-ranked— renders ranked terms/modules as either:- bars (Metascape-like horizontal bars), or
- packed circles (module-level circle packing with term circles inside).
Use rank to generate a deterministic ranked table from a run output directory.
llm-pathway-curator rank --help
# Typical workflow: point rank to a run directory and write claims_ranked.tsv
# (See --help for the exact flags supported by your installed version.)plot-ranked auto-detects claims_ranked.tsv (recommended) or falls back to audit_log.tsv
under --run-dir.
Packed circles require an extra dependency:
python -m pip install circlify
llm-pathway-curator plot-ranked \
--mode bars \
--run-dir out/demo \
--out-png out/demo/plots/ranked_bars.png \
--decision PASS \
--group-by-module \
--left-strip \
--strip-labels \
--bar-color-mode modulellm-pathway-curator plot-ranked \
--mode packed \
--run-dir out/demo \
--out-png out/demo/plots/ranked_packed.png \
--decision PASS \
--term-color-mode modulellm-pathway-curator plot-ranked \
--mode packed \
--run-dir out/demo \
--out-png out/demo/plots/ranked_packed.direction.png \
--decision PASS \
--term-color-mode directionplot-ranked assigns a single module display rank (M01, M02, ...) and a stable module color per module_id,
so bars and packed circles can be placed side-by-side without label/color drift.
Each row is one enriched term.
Required columns:
term_id,term_name,sourcestat,qval,directionevidence_genes(supporting genes; TSV uses;join)
Structured context record used for proposal and context gating, e.g.:
condition/disease,tissue,perturbation,comparison
Adapters for common tools live under src/llm_pathway_curator/adapters/.
Adapters are intentionally conservative:
- preserve evidence identity (term × genes)
- avoid destructive parsing
- keep TSV round-trips stable (contract drift is treated as a bug)
See: src/llm_pathway_curator/adapters/README.md
LLM-PathwayCurator assigns decisions by mechanical audit gates:
- FAIL: auditable violations (evidence-link drift, schema violations, contradictions, forbidden fields, etc.)
- ABSTAIN: non-specific, under-supported, or unstable under perturbations / stress tests
- PASS: survives all enabled gates at the chosen operating point (τ)
Important: the LLM (if enabled) never decides acceptance. It may propose candidates; the audit suite is the decider.
- Context swap: shuffle study context (e.g., BRCA → LUAD) to test context dependence
- Evidence dropout: randomly remove supporting genes (seeded; min_keep enforced)
- Contradiction injection (optional): introduce internally contradictory candidates to test FAIL gates
These are specification-driven perturbations intended to validate that the pipeline abstains for the right reasons, with stress-specific reason codes.
LLM-PathwayCurator is deterministic by default:
- fixed seeds (CLI + library defaults)
- pinned parsing + hashing utilities
- stable output schemas and reason codes
- run metadata persisted to
run_meta.json(and runner-levelmanifest.jsonwhen used)
Paper-side runners (e.g., paper/scripts/fig2_run_pipeline.py) orchestrate reproducible sweeps
and do not implement scientific logic; they call the library entrypoint (llm_pathway_curator.pipeline.run_pipeline).
pip install llm-pathway-curator(See PyPI project page: https://pypi.org/project/llm-pathway-curator/)
git clone https://github.com/kenflab/LLM-PathwayCurator.git
cd LLM-PathwayCurator
pip install -e .We provide an official Docker environment (Python + R + Jupyter), sufficient to run LLM-PathwayCurator and most paper figure generation.
Optionally includes Ollama for local LLM annotation (no cloud API key required).
-
Use the published image from GitHub Container Registry (GHCR).
# from the repo root (optional, for notebooks / file access) docker pull ghcr.io/kenflab/llm-pathway-curator:officialRun Jupyter:
docker run --rm -it \ -p 8888:8888 \ -v "$PWD":/work \ -e GEMINI_API_KEY \ -e OPENAI_API_KEY \ ghcr.io/kenflab/llm-pathway-curator:officialOpen Jupyter: http://localhost:8888
(Use the token printed in the container logs.)
Notes:For manuscript reproducibility, we also provide versioned tags (e.g., :0.1.0). Prefer a version tag when matching a paper release.
-
-
# from the repo root docker compose -f docker/docker-compose.yml build docker compose -f docker/docker-compose.yml upB-1.1) Open Jupyter
- http://localhost:8888
Workspace mount:
/work
B-1.2) If prompted for "Password or token"
- Get the tokenized URL from container logs:
docker compose -f docker/docker-compose.yml logs -f llm-pathway-curator
- Then either:
- open the printed URL (contains
?token=...) in your browser, or - paste the token value into the login prompt.
- open the printed URL (contains
- http://localhost:8888
Workspace mount:
-
# from the repo root docker build -f docker/Dockerfile -t llm-pathway-curator:official .
B-2.1) Run Jupyter
docker run --rm -it \ -p 8888:8888 \ -v "$PWD":/work \ -e GEMINI_API_KEY \ -e OPENAI_API_KEY \ llm-pathway-curator:officialB-2.2) Open Jupyter
- http://localhost:8888
Workspace mount:
/work
- http://localhost:8888
Workspace mount:
-
-
Use the published image from GitHub Container Registry (GHCR).
apptainer build llm-pathway-curator.sif docker://ghcr.io/kenflab/llm-pathway-curator:official
-
docker compose -f docker/docker-compose.yml build apptainer build llm-pathway-curator.sif docker-daemon://llm-pathway-curator:official
Run Jupyter (either image):
apptainer exec --cleanenv \
--bind "$PWD":/work \
llm-pathway-curator.sif \
bash -lc 'jupyter lab --ip=0.0.0.0 --port=8888 --no-browser If enabled, the LLM is confined to proposal steps and must emit schema-bounded JSON with resolvable EvidenceTable links.
Backends (example):
- Ollama:
LLMPATH_OLLAMA_HOST,LLMPATH_OLLAMA_MODEL - Gemini:
GEMINI_API_KEY - OpenAI:
OPENAI_API_KEY
Typical environment:
export LLMPATH_BACKEND="ollama" # ollama|gemini|openaiDeterministic settings are used by default (e.g., temperature=0), and runs persist
prompt/raw/meta artifacts alongside run_meta.json.
paper/ contains manuscript-facing scripts, Source Data exports, and frozen/derived artifacts (when redistributable).
paper/README.md— how to reproduce figurespaper/FIGURE_MAP.csv— canonical mapping: panel ↔ inputs ↔ scripts ↔ outputs
If you use LLM-PathwayCurator, please cite:
- bioRxiv preprint (doi: 10.64898/2026.02.18.706381)
- Zenodo archive (v0.1.0): 10.5281/zenodo.18625777
- GitHub release tag: v0.1.0
- Software RRID: RRID:SCR_027964