Before implementing any code, the first step I took was to understand the purpose and structure of a GenBank file. I learned that GenBank is a database that contains information regarding necleotide sequences and that each file holds genomic information of a specific organism. I learned about each section of the file, especially the features and origin sections.
The next step I took was researching libraries that not only allowed for the parsing of the GenBank file but also the analysis and implementation of this information onto a visual diagram. In this case the diagram had to be a circular genome map that outputs as either a JPG, PNG, or JPEG file. I found that the biopython library, a tool created for biological computation, had all the features I needed to accomplish this goal. To learn more about this library I watched a comprehensive video on YouTube and visited the official biopython website which also contained instructions and documantation that would greatly help me throughout the project.
After importing this repository into a python IDE, I followed these steps,
- Installed required libraries - These libraries included biopython to analyze the GenBank data, reportlab and bio to build the circular genome map diagram and export the map as a png/jpg file
- Parsed GenBank file - Using biopython I retrieved and stored the contents of the Genome.gb file
- Set up diagram - Created an empty diagram, circular track that would be placed onto the diagram, and a feature set that would be placed onto the track
- Built bar_display function - Bars displayed in and out of the circular track were set up as well as the labels of each gene that were retrieved from the GenBank file. Arrow format and sizing were also set
- Created text file with a list of recognition sequences - After researching enzymes in the specified organism, seven restriction enzymes and their sequences were selected to display
- Built sequence_retrieval function - contents of text file were retrieved and stored in a list
- Built sequence_placement function - retrieved enzymes were located in their recognition sites in the tomato curly stunt virus genome sequence by traversing the sequence data retrieved from the GenBank file. Enzyme sequences were set to be displayed in the genome map where they were found to be located in the organism's dna sequence
- Built output_file function - circular diagram was drawn using the installed ReportLab objects and for output, diagram was redered as a png and jpg file format
- biopython
- reportlab
- bio
Chang, Jeff, et al. Biopython Tutorial and Cookbook, 4 Sept. 2020, biopython.org/DIST/docs/tutorial/Tutorial.html#sec336.
You may not collaborate with anyone on this challenge. You are allowed to use Internet documentation. If you do use existing code (either from Github, Stack Overflow, or other sources), please cite your sources in the README.
Please follow the below instructions on how to submit your answers.
- Create a public fork of this repo and name it
ACM-Research-Coding-Challenge-S21. To fork this repo, click the button on the top right and click the "Fork" button. - Clone the fork of the repo to your computer using
git clone [the URL of your clone]. You may need to install Git for this (Google it). - Complete the Challenge based on the instructions below.
- Submit your solution by filling out this form.
Genome analysis is the identification of genomic features such as gene expression or DNA sequences in an individual's genetic makeup. A genbank file (.gb) format contains information about an individual's DNA sequence. The following dataset in Genome.gb contains a complete genome sequence of Tomato Curly Stunt Virus.
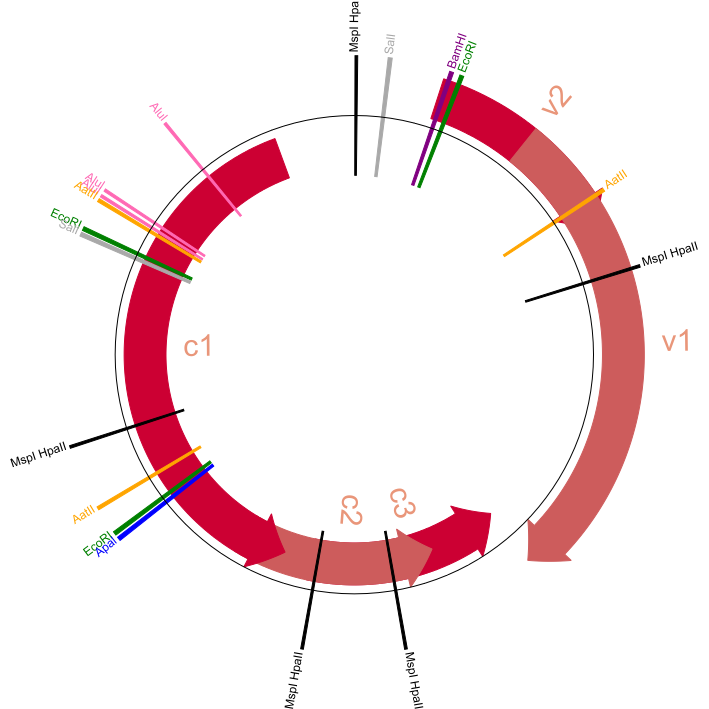
With this file, create a circular genome map and output it as a JPG/PNG/JPEG format. We're not looking for any complex maps, just be sure to highlight the features and their labels.
You may use any programming language you feel most comfortable. We recommend Python because it is the easiest to implement. You're allowed to use any library you want to implement this, just document which ones you used in this README file. Try to complete this as soon as possible.
Regardless if you can or cannot answer the question, provide a short explanation of how you got your solution or how you think it can be solved in your README.md file. However, we highly recommend giving the challenge a try, you just might learn something new!