-
Notifications
You must be signed in to change notification settings - Fork 825
Make H5MD serialize with pickle #2894
New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
Changes from all commits
5bcd6c1
d220dc5
ddfa365
b29e675
c7087b8
a40b298
94e6d63
cfcbc18
24c6090
9f28df6
d34628d
b64501f
3eb97ae
fdcd0dc
1966ee7
File filter
Filter by extension
Conversations
Jump to
Diff view
Diff view
There are no files selected for viewing
| Original file line number | Diff line number | Diff line change |
|---|---|---|
|
|
@@ -31,7 +31,7 @@ | |
| interface of the HDF5 library to improve parallel reads and writes. | ||
|
|
||
| The HDF5 library and `h5py`_ must be installed; otherwise, H5MD files | ||
| cannot be read by MDAnalysis. If `h5py`_ is not installed, a | ||
| cannot be read by MDAnalysis. If `h5py`_ is not installed, a | ||
| :exc:`RuntimeError` is raised. | ||
|
|
||
| Units | ||
|
|
@@ -94,10 +94,10 @@ | |
|
|
||
| Building parallel h5py and HDF5 on Linux | ||
| ^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^ | ||
| Building a working parallel HDF5/h5py/mpi4py environment can be | ||
| Building a working parallel HDF5/h5py/mpi4py environment can be | ||
| challenging and is often specific to your local computing resources, | ||
| e.g., the supercomputer that you're running on typically already has | ||
| its preferred MPI installation. As a starting point we provide | ||
| its preferred MPI installation. As a starting point we provide | ||
| instructions that worked in a specific, fairly generic environment. | ||
|
|
||
| These instructions successfully built parallel HDF5/h5py | ||
|
|
@@ -133,8 +133,8 @@ | |
| CC=mpicc HDF5_DIR=$HDF5_PATH python setup.py build | ||
| python setup.py install | ||
|
|
||
| If you have questions or want to share how you managed to build | ||
| parallel hdf5/h5py/mpi4py please let everyone know on the | ||
| If you have questions or want to share how you managed to build | ||
| parallel hdf5/h5py/mpi4py please let everyone know on the | ||
| `MDAnalysis forums`_. | ||
|
|
||
| .. _`H5MD`: https://nongnu.org/h5md/index.html | ||
|
|
@@ -180,6 +180,9 @@ | |
|
|
||
| .. automethod:: H5MDReader._reopen | ||
|
|
||
| .. autoclass:: H5PYPicklable | ||
| :members: | ||
|
|
||
| """ | ||
|
|
||
| import numpy as np | ||
|
|
@@ -190,6 +193,15 @@ | |
| import h5py | ||
| except ImportError: | ||
| HAS_H5PY = False | ||
|
|
||
| # Allow building documentation even if h5py is not installed | ||
| import types | ||
|
|
||
| class MockH5pyFile: | ||
| pass | ||
| h5py = types.ModuleType("h5py") | ||
| h5py.File = MockH5pyFile | ||
|
|
||
| else: | ||
| HAS_H5PY = True | ||
|
|
||
|
|
@@ -228,23 +240,23 @@ class H5MDReader(base.ReaderBase): | |
|
|
||
| See `h5md documentation <https://nongnu.org/h5md/h5md.html>`_ | ||
| for a detailed overview of the H5MD file format. | ||
| The reader attempts to convert units in the trajectory file to | ||
| the standard MDAnalysis units (:mod:`MDAnalysis.units`) if | ||
|
|
||
| The reader attempts to convert units in the trajectory file to | ||
| the standard MDAnalysis units (:mod:`MDAnalysis.units`) if | ||
| `convert_units` is set to ``True``. | ||
|
|
||
| Additional data in the *observables* group of the H5MD file are | ||
| loaded into the :attr:`Timestep.data` dictionary. | ||
|
|
||
| Only 3D-periodic boxes or no periodicity are supported; for no | ||
| periodicity, :attr:`Timestep.dimensions` will return ``None``. | ||
|
|
||
| Although H5MD can store varying numbers of particles per time step | ||
| as produced by, e.g., GCMC simulations, MDAnalysis can currently | ||
| only process a fixed number of particles per step. If the number | ||
| of particles changes a :exc:`ValueError` is raised. | ||
|
|
||
| The :class:`H5MDReader` reads .h5md files with the following | ||
| The :class:`H5MDReader` reads .h5md files with the following | ||
| HDF5 hierarchy: | ||
|
|
||
| .. code-block:: text | ||
|
|
@@ -277,21 +289,21 @@ class H5MDReader(base.ReaderBase): | |
| \-- (position) | ||
| \-- [step] <int>, gives frame | ||
| \-- [time] <float>, gives time | ||
| +-- units <str> | ||
| +-- unit <str> | ||
| \-- [value] <float>, gives numpy arrary of positions | ||
| with shape (n_atoms, 3) | ||
| +-- unit <str> | ||
| \-- (velocity) | ||
| \-- [step] <int>, gives frame | ||
| \-- [time] <float>, gives time | ||
| +-- units <str> | ||
| +-- unit <str> | ||
| \-- [value] <float>, gives numpy arrary of velocities | ||
| with shape (n_atoms, 3) | ||
| +-- unit <str> | ||
| \-- (force) | ||
| \-- [step] <int>, gives frame | ||
| \-- [time] <float>, gives time | ||
| +-- units <str> | ||
| +-- unit <str> | ||
| \-- [value] <float>, gives numpy arrary of forces | ||
| with shape (n_atoms, 3) | ||
| +-- unit <str> | ||
|
|
@@ -557,13 +569,15 @@ def open_trajectory(self): | |
| if self._comm is not None: | ||
| # can only pass comm argument to h5py.File if driver='mpio' | ||
| assert self._driver == 'mpio' | ||
| self._file = h5py.File(self.filename, 'r', # pragma: no cover | ||
| driver=self._driver, | ||
| comm=self._comm) | ||
| self._file = H5PYPicklable(name=self.filename, # pragma: no cover | ||
| mode='r', | ||
| driver=self._driver, | ||
| comm=self._comm) | ||
| elif self._driver is not None: | ||
| self._file = h5py.File(self.filename, 'r', driver=self._driver) | ||
| self._file = H5PYPicklable(name=self.filename, mode='r', | ||
| driver=self._driver) | ||
| else: | ||
| self._file = h5py.File(self.filename, 'r') | ||
| self._file = H5PYPicklable(name=self.filename, mode='r') | ||
| # pulls first key out of 'particles' | ||
| # allows for arbitrary name of group1 in 'particles' | ||
| self._particle_group = self._file['particles'][ | ||
|
|
@@ -727,3 +741,88 @@ def has_forces(self): | |
| @has_forces.setter | ||
| def has_forces(self, value: bool): | ||
| self._has['force'] = value | ||
|
|
||
| def __getstate__(self): | ||
| state = self.__dict__.copy() | ||
| del state['_particle_group'] | ||
| return state | ||
|
|
||
| def __setstate__(self, state): | ||
| self.__dict__ = state | ||
| self._particle_group = self._file['particles'][ | ||
| list(self._file['particles'])[0]] | ||
| self[self.ts.frame] | ||
|
|
||
|
|
||
| class H5PYPicklable(h5py.File): | ||
| """H5PY file object (read-only) that can be pickled. | ||
|
|
||
| This class provides a file-like object (as returned by | ||
| :class:`h5py.File`) that, | ||
| unlike standard Python file objects, | ||
| can be pickled. Only read mode is supported. | ||
|
|
||
| When the file is pickled, filename, mode, driver, and comm of | ||
| :class:`h5py.File` in the file are saved. On unpickling, the file | ||
| is opened by filename, mode, driver. This means that for a successful | ||
| unpickle, the original file still has to be accessible with its filename. | ||
|
|
||
| Parameters | ||
| ---------- | ||
| filename : str or file-like | ||
| a filename given a text or byte string. | ||
| driver : str (optional) | ||
| H5PY file driver used to open H5MD file | ||
|
|
||
| Example | ||
| ------- | ||
| :: | ||
|
|
||
| f = H5PYPicklable('filename', 'r') | ||
| print(f['particles/trajectory/position/value'][0]) | ||
| f.close() | ||
|
|
||
| can also be used as context manager:: | ||
|
|
||
| with H5PYPicklable('filename', 'r'): | ||
| print(f['particles/trajectory/position/value'][0]) | ||
|
|
||
| Note | ||
| ---- | ||
| Pickling of an `h5py.File` opened with `driver="mpio"` and an MPI | ||
| communicator is currently not supported | ||
|
|
||
| See Also | ||
| --------- | ||
| :class:`MDAnalysis.lib.picklable_file_io.FileIOPicklable` | ||
| :class:`MDAnalysis.lib.picklable_file_io.BufferIOPicklable` | ||
| :class:`MDAnalysis.lib.picklable_file_io.TextIOPicklable` | ||
| :class:`MDAnalysis.lib.picklable_file_io.GzipPicklable` | ||
| :class:`MDAnalysis.lib.picklable_file_io.BZ2Picklable` | ||
|
|
||
|
|
||
| .. versionadded:: 2.0.0 | ||
| """ | ||
|
|
||
| def __getstate__(self): | ||
| driver = self.driver | ||
| # Current issues: Need a way to retrieve MPI communicator object | ||
| # from self and pickle MPI.Comm object. Parallel driver is excluded | ||
| # from test because h5py calls for an MPI configuration when driver is | ||
| # 'mpio', so this will need to be patched in the test function. | ||
| if driver == 'mpio': # pragma: no cover | ||
| raise TypeError("Parallel pickling of `h5py.File` with" # pragma: no cover | ||
| " 'mpio' driver is currently not supported.") | ||
|
|
||
| return {'name': self.filename, | ||
|
Member
There was a problem hiding this comment. Choose a reason for hiding this commentThe reason will be displayed to describe this comment to others. Learn more. I am confused: If the
Member
There was a problem hiding this comment. Choose a reason for hiding this commentThe reason will be displayed to describe this comment to others. Learn more. I see that you had it included in b29e675 but then removed again. Did you test that a from mpi4py import MPI
import pickle
comm = MPI.COMM_WORLD
comm_p = pickle.loads(pickle.dumps(comm))
assert comm == comm_por something similar?
Member
Author
There was a problem hiding this comment. Choose a reason for hiding this commentThe reason will be displayed to describe this comment to others. Learn more.
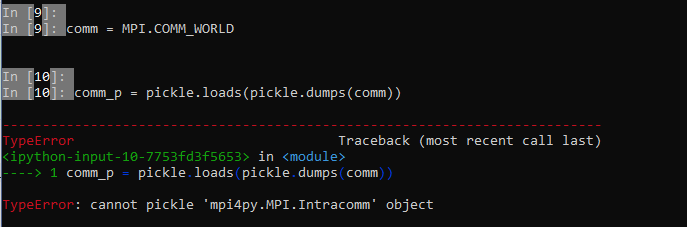
Contributor
There was a problem hiding this comment. Choose a reason for hiding this commentThe reason will be displayed to describe this comment to others. Learn more. I don’t have a working mpi4py in my desktop. But it seems that it cannot be pickled natively. |
||
| 'mode': self.mode, | ||
| 'driver': driver} | ||
|
|
||
| def __setstate__(self, state): | ||
| self.__init__(name=state['name'], | ||
| mode=state['mode'], | ||
| driver=state['driver']) | ||
|
|
||
| def __getnewargs__(self): | ||
| """Override the h5py getnewargs to skip its error message""" | ||
| return () | ||
Uh oh!
There was an error while loading. Please reload this page.